Request Demo
Last update 14 Jun 2025
RBD-S-Dps
Last update 14 Jun 2025
Overview
Basic Info
Drug Type Prophylactic vaccine |
Synonyms- |
Target |
Action inhibitors |
Mechanism SARS-CoV-2 S protein inhibitors(SARS-CoV-2 S protein inhibitors) |
Therapeutic Areas |
Active Indication- |
Inactive Indication |
Originator Organization |
Active Organization- |
Inactive Organization |
License Organization- |
Drug Highest PhasePendingPreclinical |
First Approval Date- |
Regulation- |
R&D Status
10 top R&D records. to view more data
Login
| Indication | Highest Phase | Country/Location | Organization | Date |
|---|---|---|---|---|
| Coronavirus Infections | Preclinical | Global | 01 Sep 2021 |
Login to view more data
Clinical Result
Clinical Result
Indication
Phase
Evaluation
View All Results
Login to view more data
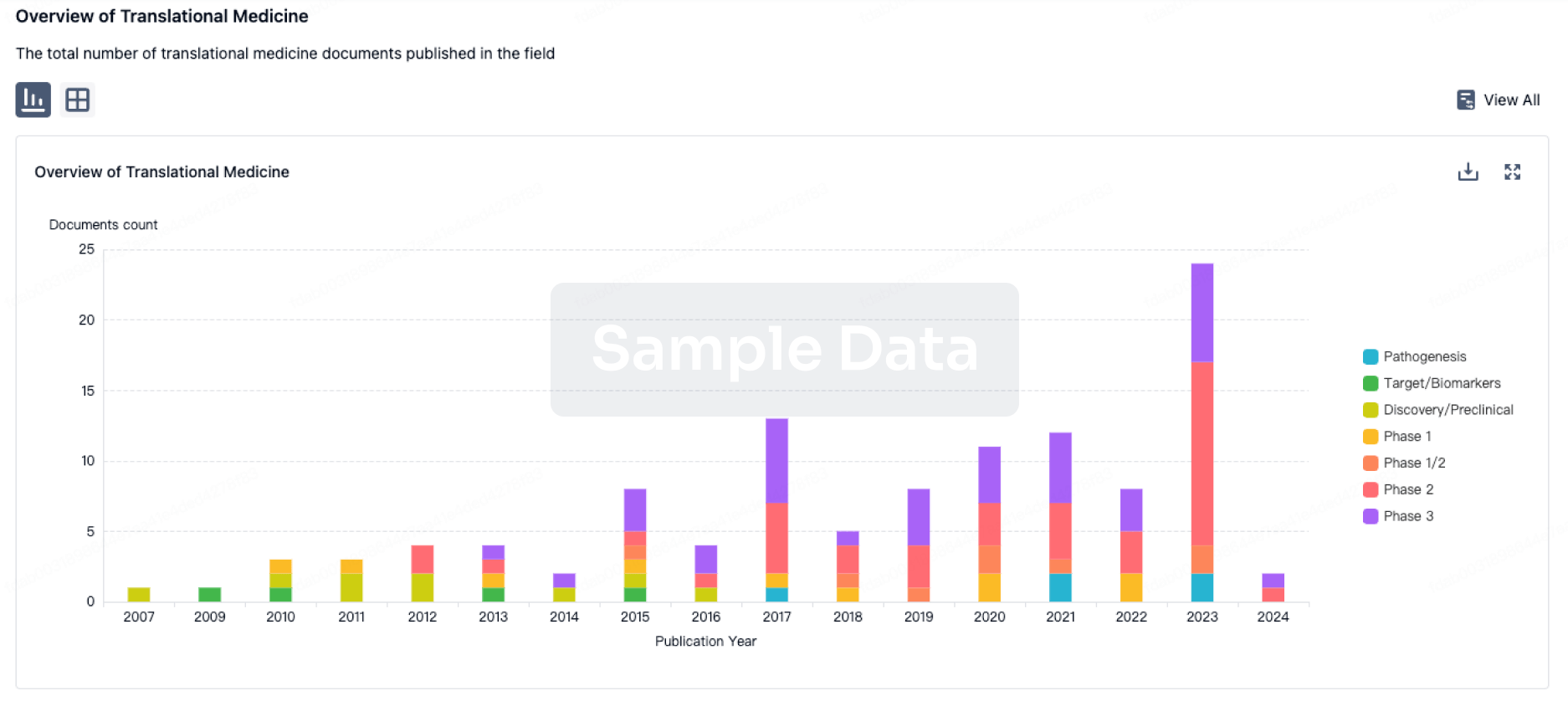
Translational Medicine
Boost your research with our translational medicine data.
login
or
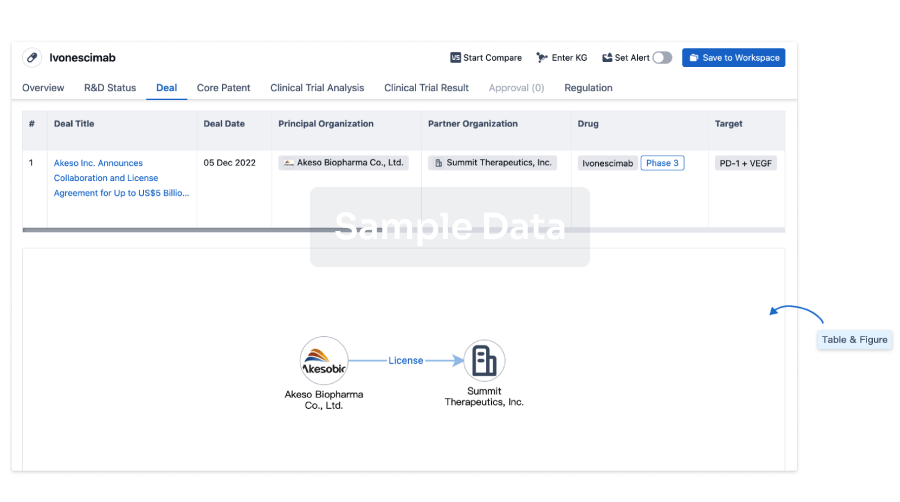
Deal
Boost your decision using our deal data.
login
or
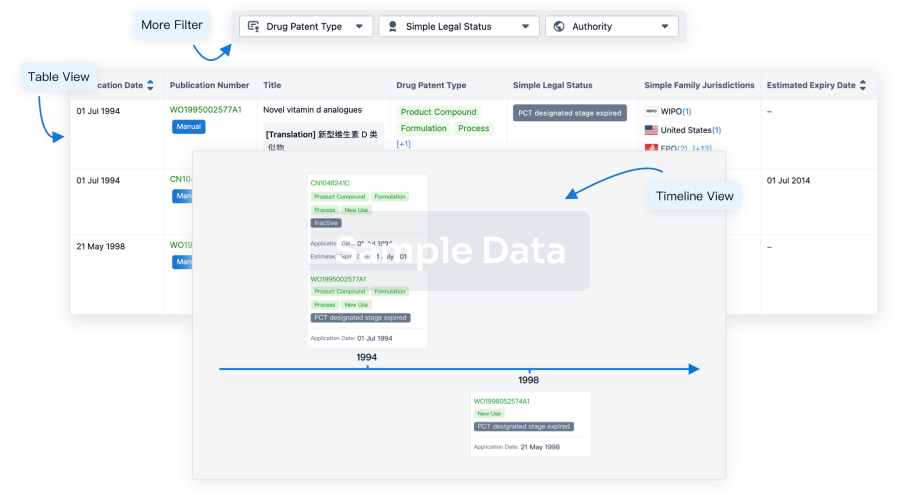
Core Patent
Boost your research with our Core Patent data.
login
or
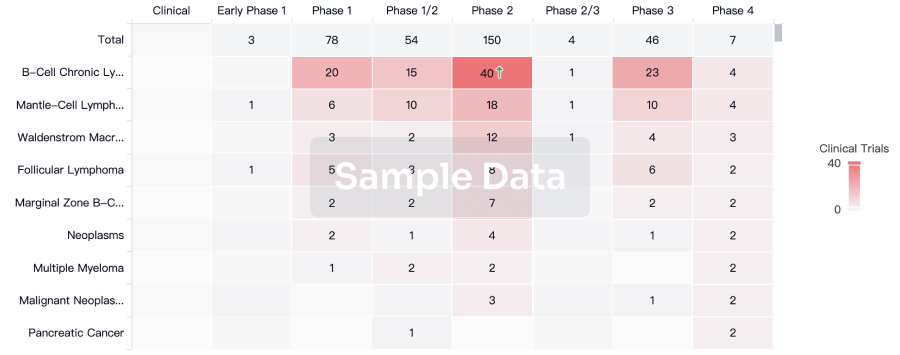
Clinical Trial
Identify the latest clinical trials across global registries.
login
or
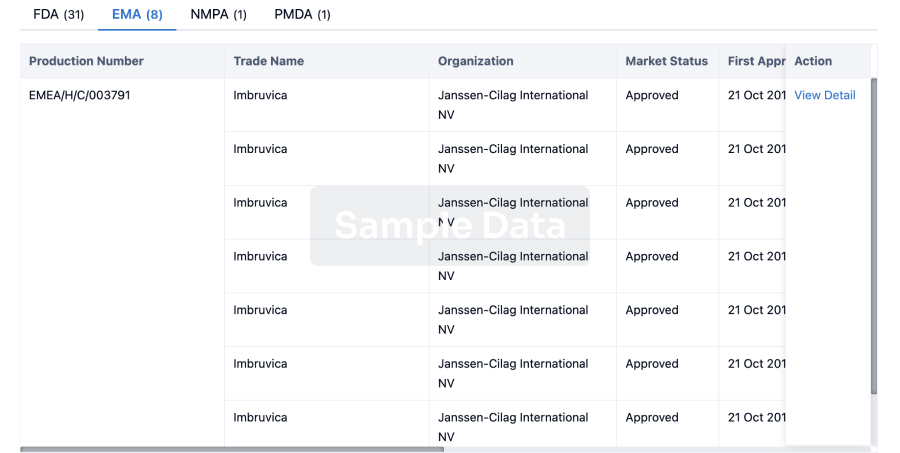
Approval
Accelerate your research with the latest regulatory approval information.
login
or
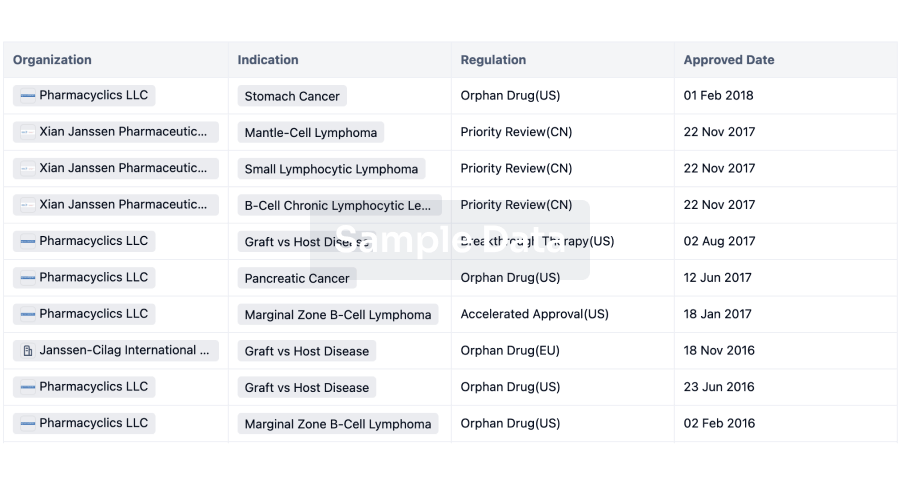
Regulation
Understand key drug designations in just a few clicks with Synapse.
login
or
AI Agents Built for Biopharma Breakthroughs
Accelerate discovery. Empower decisions. Transform outcomes.
Get started for free today!
Accelerate Strategic R&D decision making with Synapse, PatSnap’s AI-powered Connected Innovation Intelligence Platform Built for Life Sciences Professionals.
Start your data trial now!
Synapse data is also accessible to external entities via APIs or data packages. Empower better decisions with the latest in pharmaceutical intelligence.
Bio
Bio Sequences Search & Analysis
Sign up for free
Chemical
Chemical Structures Search & Analysis
Sign up for free
