Request Demo
Last update 20 Jun 2025

Clalit Research Institute
Last update 20 Jun 2025
Overview
Related
2
Clinical Trials associated with Clalit Research InstituteNCT04523324
Randomized Comparison of the Immunogenicity of Recombinant and Egg-based Influenza Vaccines Among Healthcare Personnel in Israel
This randomized, open-label, active-controlled trial will assess humoral immune responses to a single dose of 2019-20 recombinant hemagglutinin quadrivalent influenza vaccines (RIV4) compared with standard egg-based unadjuvanted quadrivalent influenza vaccines (IIV4) among healthcare personnel (HCP) vaccinated during the previous 2018-19 season with IIV4. The trial will be conducted at two hospital sites in Israel during the 2019-20 influenza season among HCP who were enrolled in the Study of Healthcare Personnel with Influenza and other Respiratory Viruses in Israel (SHIRI).
Start Date31 Oct 2019 |
Sponsor / Collaborator |
NCT03331991
Prevention of Influenza and Other Wintertime Respiratory Viruses Among Healthcare Professionals in Israel
The purpose of the study on the Prevention of Influenza and Other Wintertime Respiratory Viruses among Healthcare Professionals in Israel Effectiveness of Influenza Vaccine in Preventing Influenza Virus Infection, Missed Work, and Patient Exposure: A Prospective Cohort Study of Healthcare Personnel (to be called the Healthcare Personnel or HCP study throughout this Data Security Plan) is to investigate vaccine effectiveness and respiratory illness among healthcare personnel (HCP). This will help to better understand the factors that influence influenza vaccination choice, individual vaccine response, and whether or not the influenza vaccine helps to prevent influenza in HCP.
Start Date28 Sep 2016 |
Sponsor / Collaborator ABT Associates [+2] |
100 Clinical Results associated with Clalit Research Institute
Login to view more data
0 Patents (Medical) associated with Clalit Research Institute
Login to view more data
173
Literatures (Medical) associated with Clalit Research Institute04 Apr 2025·Journal of Crohns & Colitis
GLP-1 Analog Use is Associated With Improved Disease Course in Inflammatory Bowel Disease: A Report from the Epi-IIRN
Article
Author: Matz, Eran ; Turner, Dan ; Ghersin, Itai ; Loewenberg Weisband, Yiska ; Shlon, Dima ; Ben-Tov, Amir ; Gorelik, Yuri ; Dotan, Iris ; Zacay, Galia ; Bar-Yoseph, Haggai ; Lujan, Rona
Abstract:
Background and Aims:
The growing use of glucagon-like peptide 1 (GLP-1) analogs for type 2 diabetes mellitus (DM2) and obesity necessitates studies about their use in patients with inflammatory bowel diseases (IBD).
Methods:
Data on patients with DM2 were retrieved from an Israeli nationwide cohort of patients with IBD (epi-IIRN), recording GLP-1 analog exposure for at least 6 months. The primary outcome was poor disease outcomes (ie, composite of steroid dependence, initiation of advanced IBD therapy, hospitalization, surgery, or death). Cox proportional hazard models with time-varying covariables were used to assess the impact of GLP-1 use on outcomes during follow-up.
Results:
We included 3737 patients (24 338 patient-years) with IBD and DM2 [50.4% ulcerative colitis (UC)], of whom 633 were treated with GLP-1 analogs. Accounting for demographics IBD/DM2 related variables, medication use, and laboratory measurements, GLP-1 analog use was associated with reduced composite outcome in the full cohort (adjusted hazard ratio [aHR] 0.74, 95% confidence interval [CI] 0.62-0.89) and in each subtype [UC (aHR 0.71, 95% CI 0.52-0.96) and Crohn’s disease (aHR 0.78, 95% CI 0.62-0.99)]. Similar trends were seen in multivariate analyses of each individual outcome, although only hospitalization was significant (aHR 0.74, 95% CI 0.61-0.91). The protective effect of GLP-1 analogs was seen in patients with obesity (aHR 0.61, 95% CI 0.50-0.77), but not in non-obese (aHR 0.94, 95% CI 0.67-1.31).
Conclusions:
GLP-1 analogs are associated with improved outcomes in IBD, specifically in patients with obesity. The mechanisms of these effects require further investigation as well as their role in patients without DM2.
01 Apr 2025·Journal of Genetic Counseling
Lessons learned from BRCA1 /2 screening in Israel: A cross‐sectional survey comparing experiences and communication
Article
Author: Sagi‐Dain, Lena ; Greenberg, Rotem ; Singer, Amihood ; Echar, Moran
Abstract:
This study aimed to evaluate the perceived quality of pre‐ and post‐test explanations given to women carrying BRCA1/2 variants, and to compare these outcomes between two cohorts: female BRCA1/2 carriers identified through self‐reported population‐based screening (the screening group), in comparison to self‐reported formal pre‐test genetic counseling due to personal or familial cancer history (genetic counseling group). This cross‐sectional survey of female BRCA1/2 carriers employed an anonymous questionnaire distributed through the “Good Genes – a support and information group for BRCA carriers” association from January to March 2023. Main evaluated outcomes included the perceived quality of pre‐ and post‐test explanations, first analyzed in the overall cohort, and then compared between the 110 respondents in the screening group, to 444 women in the counseling group. In the screening group, 45.5% rated the perceived quality of pre‐test explanations as unsatisfactory, compared to 27.4% in the genetic counseling group (p = 0.0005). In terms of result delivery, the screening group reported higher instances of inappropriate timing (61.8% vs. 40.3%, p < 0.0001), suboptimal mode of delivery (55.5% vs. 37.5%, p = 0.0008) and suboptimal perceived quality in post‐test explanations (51.4% vs. 33.9%, p = 0.0006), as well as elevated stress levels (74.3% vs. 64.3%, p = 0.043). In the screening group, 21.5% of the women reported that the results were communicated by phone, letter, or online notice, compared to 17.2% in the counseling group, a non‐statistically significant difference. A logistic regression model controlling for timing and mode of delivery showed that both timing (β = 0.46, p < 0.001) and mode of delivery (β = 0.39, p < 0.001) remained significant predictors of dissatisfaction of post‐test counseling. The findings of this survey underscore the pressing need for enhancements in pre‐test explanation, as well as the post‐test counseling for positive results, especially within the realm of BRCA screening.
01 Mar 2025·Clinical Gastroenterology and Hepatology
Risk of Age-related and Disease-related Complications and Mortality in Elderly-onset Inflammatory Bowel Disease - A Population-based Study
Article
Author: Lebedenko, Boris ; Ben Hur, Dana ; Waterman, Matti ; Turner, Dan ; Dotan, Iris ; Haklai, Ziona ; Pinto, Gabriel D ; Lederman, Natan ; Issaschar, Guy ; Matz, Eran ; Moshe, Ran ; Ben-Tov, Amir ; Loewenberg Weisband, Yiska ; Lujan, Rona
BACKGROUND & AIMS:
In this nationwide cohort from Israel (Epi-IIRN), we aimed to characterize risks for age-related complications, mortality, and inflammatory bowel disease (IBD)-related surgeries in patients with elderly-onset IBD (EO-IBD).
METHODS:
Data of patients with EO-IBD (≥65 years) diagnosed during 2005 to 2020 were retrieved from the epi-IIRN database. Patients with EO-IBD were compared with 3 age-, sex-, and district-matched non-IBD individuals, for age-related outcomes. Patients with incident EO-IBD were matched to 4 adult-onset (AO) IBD (≥18-65 years) by IBD subtype, sex, and district. Cumulative incidence functions were calculated to estimate event probabilities over time, accounting for death as a competing risk. Proportional subdistribution hazards models were used to assess predictors of medication use, surgery, and complications.
RESULTS:
Of 2826 EO-IBD cases, 2162 had 3 matched non-IBD controls. Mortality rates per 1000 person-years (PY) were similar in EO-IBD and non-IBD controls (292.32; 95% confidence interval [CI], 273.53-311.85 vs 291.24; 95% CI, 280.31-302.42, respectively) as were mortality causes and risk for pneumonia (adjusted hazard rate [aHR], 1.04; 95% CI, 0.84-1.29), fractures (aHR, 1.03; 95% CI, 0.82-1.29), bacteremia (aHR, 2.16; 95% CI, 0.87-5.40), and thromboembolism (aHR, 0.58; 95% CI, 0.27-1.23). When matching 2826 patients with EO-IBD to 11,304 patients with AO-IBD, the EO-IBD group had lower exposure to thiopurines (aHR, 0.44; 95% CI, 0.39-0.49) and anti-tumor necrosis factor (TNF) (aHR, 0.37; 95% CI, 0.32-0.42) and higher risk for abdominal surgery (aHR, 1.23; 95% CI, 1.04-1.46) in Crohn's disease [CD]; aHR, 1.51; 95% CI, 2.04-3.08 in ulcerative colitis [UC], respectively) but lower perianal surgery risk (hazard ratio [HR], 0.27; 95% CI, 0.16-0.47) in CD. The calculated frequencies of repeat perianal and abdominal surgery in the EO-CD and AO-CD groups at 3 years were 7.1% and 36%, respectively, and 29% and 21%, respectively.
CONCLUSIONS:
Compared with non-IBD elderly, patients with EO-IBD have similar risks for death and complications. Compared with AO-IBD, patients with EO-IBD are at higher risk for abdominal surgery, but not for perianal surgery.
15
News (Medical) associated with Clalit Research Institute21 Feb 2024
The study assessed the sensitivity, specificity, and usability of PressureSafe against the standard of care in the detection of early-stage pressure injuries Stage 1 / suspected deep tissue injuries before skin breakage
IR-MED reports positive efficacy data for its PressureSafe device. (Credit: Nappy on Unsplash)
IR-MED, a developer of a spectrographic analysis technology platform, said that its PressureSafe device has shown 92% efficacy in early, non-invasive, skin colour agnostic detection of pressure injuries.
PressureSafe is a non-invasive handheld optical monitoring device. It leverages infrared optical spectroscopy and an artificial intelligence (AI)-based algorithm to detect early-stage pressure injuries for all skin tones.
The single arm, bi-centre study of the device was conducted at two medical centres owned by Clalit, a health maintenance organisation.
The 14-day efficacy portion of the study was intended to assess the sensitivity, specificity, and usability of PressureSafe.
It was compared against the standard of care in the detection of early-stage pressure injuries Stage 1 / suspected deep tissue injuries (sDTI) before skin breakage.
IR-MED assessed 38 patients at high risk of pressure injury development. On 154 body sites, 924 scans were performed in total.
The highly favourable proof of efficacy data showed that PressureSafe was able to identify Stage 1 / sDTI pressure injuries with 92% sensitivity and 88% specificity.
The study also assessed device calibration and validation in addition to safety. A total of 1,493 scans yielded no safety signals, out of the 66 patients whose data was acquired for safety study.
Additionally, the rate of incidence of pressure injuries diminished by 50% during the study period.
The medical device firm concluded that PressureSafe is a useful, effective, and safe decision support device for the early diagnosis of pressure injuries based on these results.
IR-MED CEO Tzur Di-Cori said: “This robust and impressive data from our collaborative study with Clalit comes at an ideal time as we prepare to enter the US market with PressureSafe.
“We believe the 92% efficacy findings in real-world data settings at two world-class hospitals will be a big factor in driving adoption in the US.”
The handheld optical monitoring device is currently in usability trials at various medical centres.
Clinical Result
07 Sep 2023
LINKÖPING, Sweden, Sept. 7, 2023 /PRNewswire/ --
International medical imaging IT and cybersecurity company Sectra's (STO: SECT B)
solution for digital pathology will be utilized by Israel's largest healthcare provider—Clalit Health Services. The solution was sold through Sectra's distribution partner in Israel, VaRay Oncology Systems. It aims to simplify the work of Clalit's pathologists in reviewing and collaborating on cases, leading to more precise cancer diagnostics.
"We are looking forward to implement Sectra's digital pathology solution," says Dr. Orly Weinstein, Deputy Director General, Head of Hospital Division, Clalit Health Services. "It will definitely enhance diagnostic capabilities, efficiency, collaboration and research potential. Digitizing pathology will allow us to take advantage of our network's capacities, as well as improve our reporting time and by doing so we will provide diagnosis faster and in a more efficient way".
The solution was sold by Sectra's distribution partner in Israel, VaRay Oncology Systems, to Clalit Health Services, a healthcare network encompassing nine sites and serving over 4.9 million members.
The contract, signed in August 2023, includes Sectra's digital pathology solution and cloud-based storage service using Microsoft Azure. This addition will enhance the capabilities of their pathologists by introducing a digital solution alongside traditional microscopes, enabling them to collaborate and review cases in a way that was previously impossible. The digital workflow provides instant access to digital images of tissue samples, allowing for remote access as needed. This eliminates the need to rely on physical glass slides viewed through microscopes.
Sectra's digital pathology solution is part of its enterprise imaging offering. It provides a unified strategy for all imaging needs while lowering operational costs. The scalable and modular solution, with a VNA at its core, allows healthcare providers to grow from ology to ology and from enterprise to enterprise. Visit Sectra's website to read more about Sectra and why it's top-ranked in "Best in KLAS".
About Sectra
Sectra contributes to a healthier and safer society by assisting health systems throughout the world to enhance the efficiency of care, and authorities and defense forces in Europe to protect society's most sensitive information. The company, founded in 1978, is headquartered in Linköping, Sweden, with direct sales in 19 countries, and distribution partners worldwide. Sales in the 2022/2023 fiscal year totaled SEK 2,351 million. The Sectra share is quoted on the Nasdaq Stockholm exchange. For more information, visit Sectra's website.
About Clalit
Clalit is the largest provider of public health services in Israel. For over 100 years, Clalit has been at the forefront of medical care and health innovations in Israel. Clalit has over 4.9 million customers, which are 53% of market share and is the largest civil employer in the country with 49,000 employees. Clalit owns and manages over 1,600 Primary Clinics and 14 hospitals (30% of Israel's hospital beds) spread all over the country. Clalit's Hospitals provide comprehensive healthcare services to a large portion of Israel's population. They focus on preventive care, research and innovation, promoting overall health and wellness. Additionally, Clalit's hospitals extensive network and integrated electronic health record system enhances patient care coordination and efficiency.
For further information, please contact:
Dr. Torbjörn Kronander,
CEO and President Sectra AB,
46 (0) 705 23 52 27
Marie Ekström Trägårdh,
Executive Vice President Sectra AB and President Sectra Imaging IT Solutions,
46 (0)708 23 56 10
The following files are available for download:
SOURCE Sectra
31 May 2023
Israel's largest health services provider partners with Cloudera to use a data lake in its private cloud to manage and analyze data from over half the country's population in real-time for quicker and better-informed decision-making, improving the quality of patient care.
TEL AVIV, Israel, May 31, 2023 /PRNewswire/ -- Cloudera, the hybrid data company, today announced its collaboration with Clalit Health Services, the largest of Israel's four state-mandated health service organizations. The health service provider is set to harness the Cloudera Data Platform (CDP) to provide high-quality, easy-to-access and around-the-clock innovative medical services to its patients, wherever they are.
Clalit will work with Cloudera to improve patient care for its 4.8 million members by building a next generation data lake on CDP. The secure platform integrates into the health provider's existing system and allows for real-time analysis and faster processing times for insight-based decision-making, enabling Clalit to enhance its streaming analytics and handle the ever-increasing volumes of data coming in.
Liora Shechter, VP Digital & Technology, Clalit Health Services, said, "We will use Cloudera Data Platform to apply real time interfaces, artificial intelligence and machine learning to further enhance patient care. This will include enhancing our big data and artificial intelligence-based C-Pi platform, which offers a powerful set of tools based on AI to identify high-risk patients who require proactive intervention. By mapping detailed clinical pathways based on the most up-to-date medical guidelines, cross-referenced with the vast amount of digital data in each patient's medical record, we can provide proactive treatment recommendations during routine medical encounters. This helps clinicians identify gaps in patient care, even if they arrive due to other complaints. CDP will be instrumental in powering C-Pi so it can provide in-depth support for the decision-making process and help clinicians stay ahead of the curve. With CDP, we'll be able to proactively reach patients who need care, even before the patients know the need care."
The health service provider will improve patient care by analyzing streams of data from electronic health records (EHRs), wearable devices and remote monitoring devices using Cloudera's Apache NiFi software and CDP. This will enable Clalit to remotely monitor patients' vital signs and recognize patterns of activity that may be indicative of a health condition, helping to identify potential problems and intervene before they become more serious. Furthermore, with enhanced capability to automate the flow of structured and unstructured data between different systems and databases, Clalit can share and access information more efficiently and coordinate care more effectively, resulting in improved patient outcomes.
Oran Sharon, Regional Vice President, Cloudera, comments, "Clalit is recognized for its innovative approach to healthcare delivery and its use of technology to improve the quality and efficiency of its services. We are delighted to support Clalit in using data to transform its business to provide even better services based on CDP."
Clalit is the second largest employer in Israel, with over 45,000 employees, including 10,000 physicians and 13,000 nurses. As such, the organization must have a robust and secure data platform to effectively manage and analyze the vast amounts of data it generates so it can provide customized and high-quality healthcare to the millions of patients that rely on it.
About Cloudera
At Cloudera, we believe data can make what is impossible today, possible tomorrow. Cloudera taught the world the value of big data, creating an industry and ecosystem powered by the relentless innovation of the open source community. We empower our customers, leaders in their industries, to transform complex data into clear and actionable insights. Through our hybrid data cloud platform, organizations are able to build their data-driven future by getting data - no matter where it resides - into the hands of those that need it. Learn more at Cloudera.com.
Cloudera and associated marks are trademarks or registered trademarks of Cloudera, Inc. All other company and product names are trademarks of their respective owners.
About Clalit Health Services
Clalit, the largest and leading HMO in Israel, provides medical services to more than 52% of the Israeli population, with a wide range of medical services: outpatient clinics throughout the country, where there are women's health centers, pediatric health centers, and specialized medical centers. In addition, Clalit manages 14 public hospitals, and provides an additional set of services that include the network of dental clinics Clalit Smile, Clalit Aesthetics, Clalit Complementary Medicine, and 'Mor' Medical Institutes and Services. Clalit works to prevent disease and promote a healthy lifestyle through offering quality medical services, and advanced on-line services that provide the ability to receive medical services from anywhere, and at any time.
For more information:
SOURCE Cloudera, Inc.
100 Deals associated with Clalit Research Institute
Login to view more data
100 Translational Medicine associated with Clalit Research Institute
Login to view more data
Corporation Tree
Boost your research with our corporation tree data.
login
or
Pipeline
Pipeline Snapshot as of 07 Nov 2025
No data posted
Login to keep update
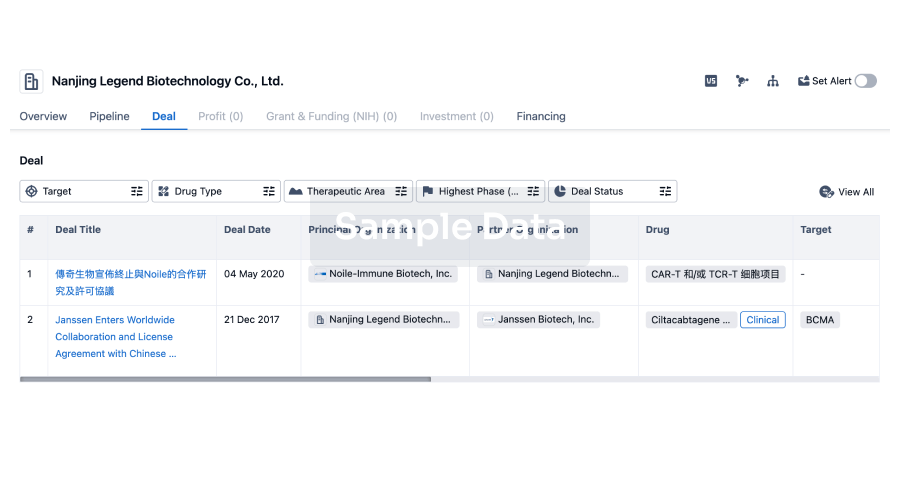
Deal
Boost your decision using our deal data.
login
or
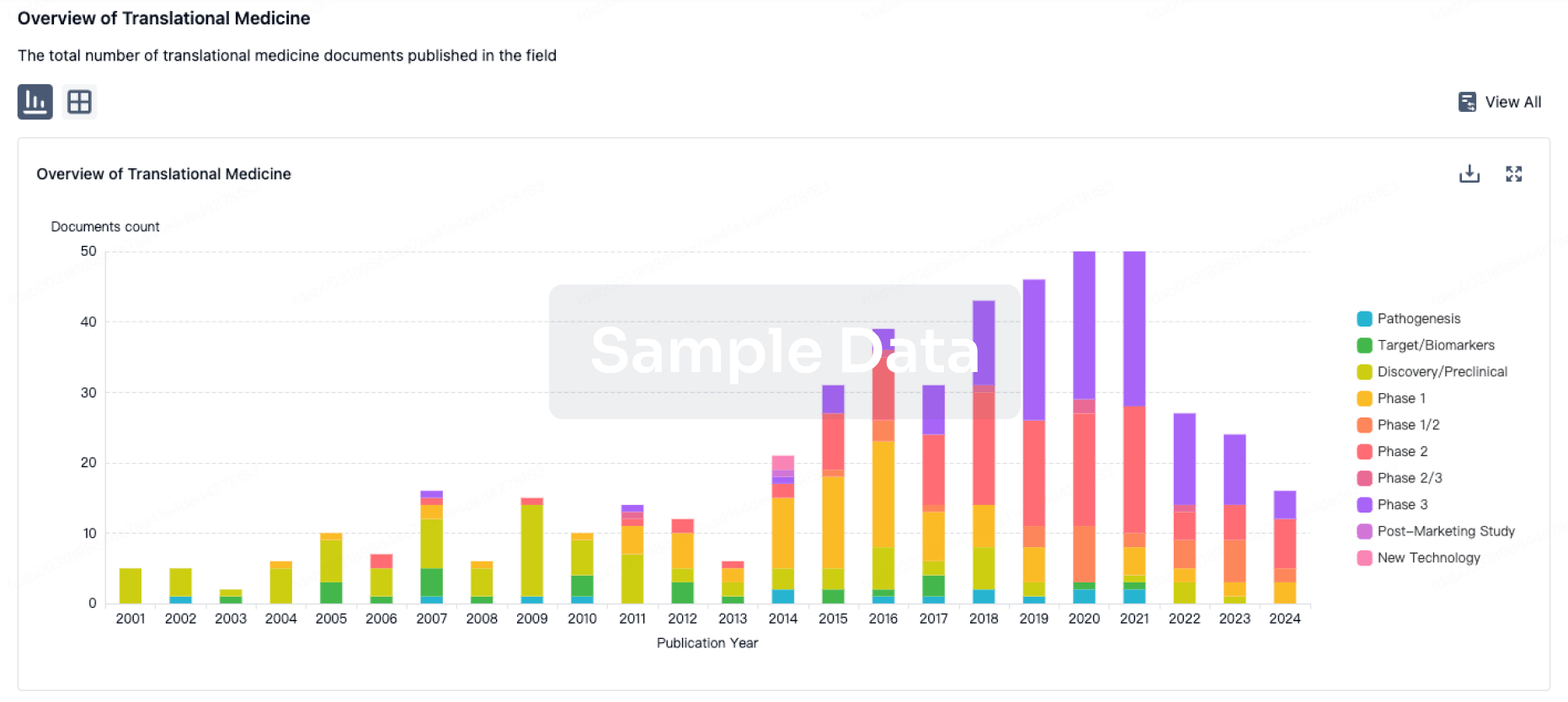
Translational Medicine
Boost your research with our translational medicine data.
login
or
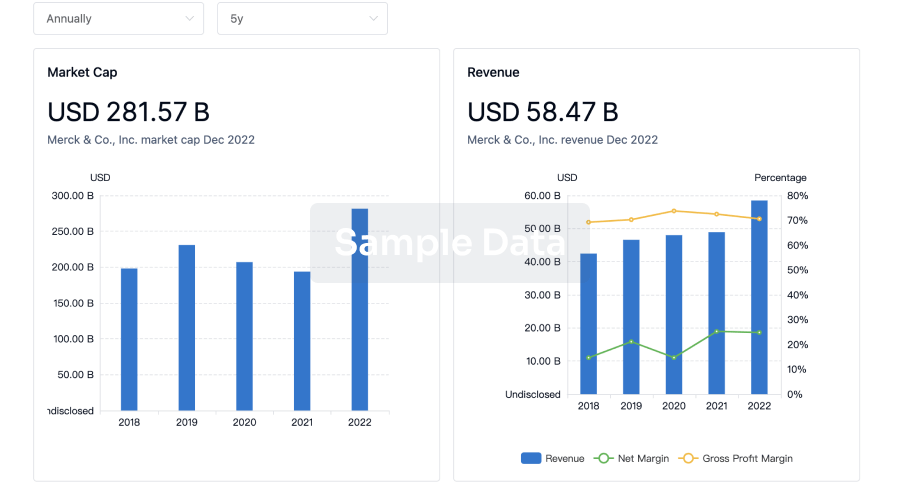
Profit
Explore the financial positions of over 360K organizations with Synapse.
login
or
Grant & Funding(NIH)
Access more than 2 million grant and funding information to elevate your research journey.
login
or
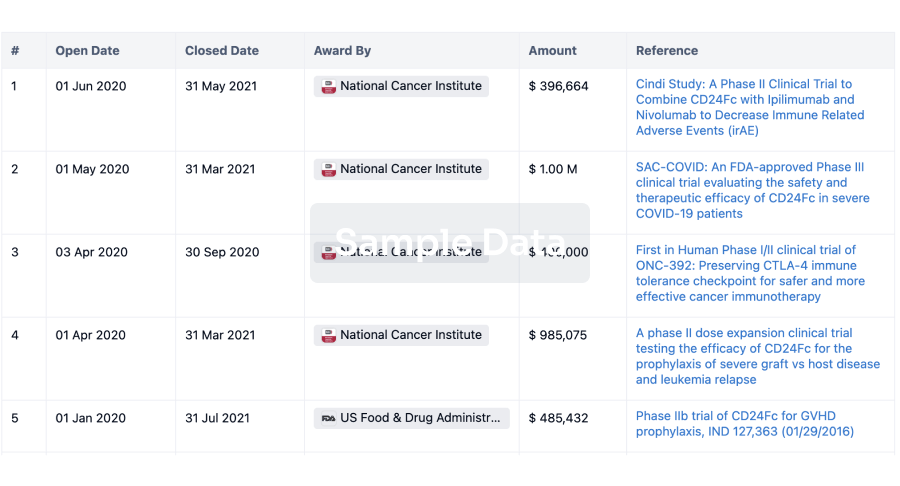
Investment
Gain insights on the latest company investments from start-ups to established corporations.
login
or
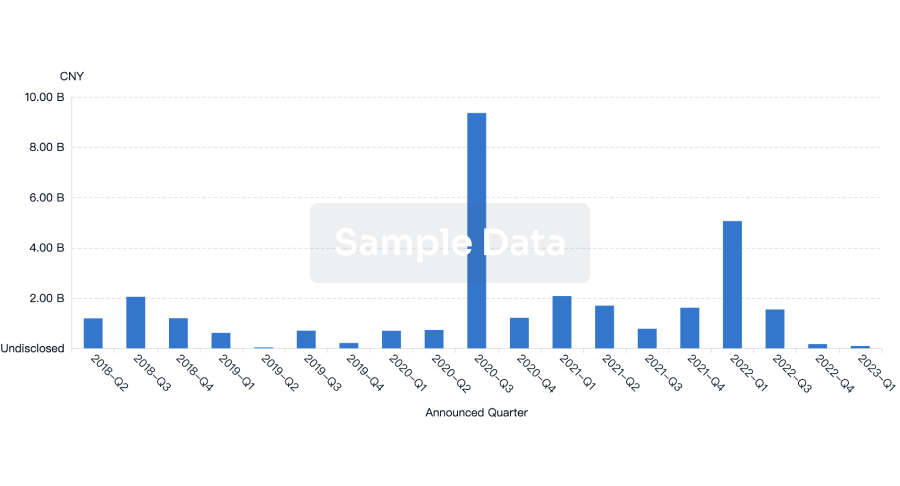
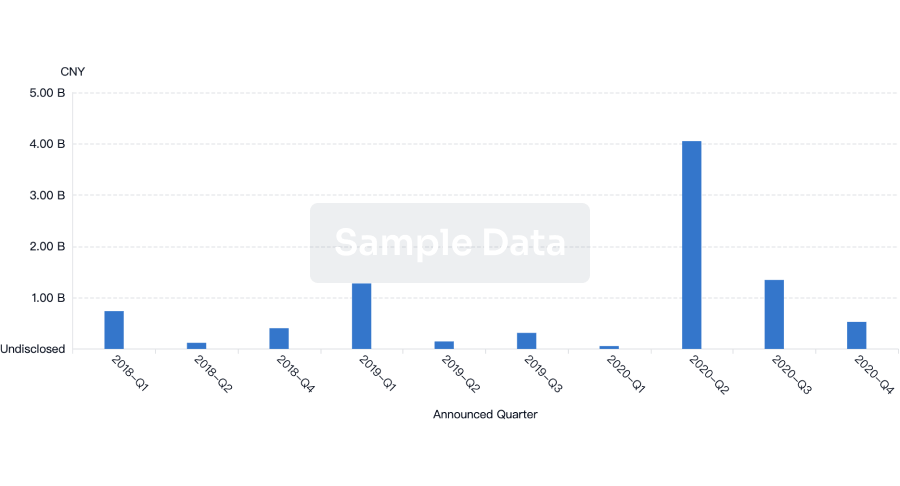
Financing
Unearth financing trends to validate and advance investment opportunities.
login
or
AI Agents Built for Biopharma Breakthroughs
Accelerate discovery. Empower decisions. Transform outcomes.
Get started for free today!
Accelerate Strategic R&D decision making with Synapse, PatSnap’s AI-powered Connected Innovation Intelligence Platform Built for Life Sciences Professionals.
Start your data trial now!
Synapse data is also accessible to external entities via APIs or data packages. Empower better decisions with the latest in pharmaceutical intelligence.
Bio
Bio Sequences Search & Analysis
Sign up for free
Chemical
Chemical Structures Search & Analysis
Sign up for free
