Request Demo
Last update 13 Nov 2024
ITGB8
Last update 13 Nov 2024
Basic Info
Related Targets |
Related
2
Drugs associated with ITGB8WO2024056668
Patent MiningTarget |
Mechanism- |
Active Org. |
Originator Org. |
Active Indication |
Inactive Indication- |
Drug Highest PhaseDiscovery |
First Approval Ctry. / Loc.- |
First Approval Date- |
WO2023244547
Patent MiningTarget |
Mechanism- |
Active Org. |
Originator Org. |
Active Indication |
Inactive Indication- |
Drug Highest PhaseDiscovery |
First Approval Ctry. / Loc.- |
First Approval Date- |
1
Clinical Trials associated with ITGB8NCT04365218
A Phase I Randomized, Blinded, Placebo-controlled Study to Evaluate the Safety and Pharmacokinetics of MEDI8367 Administered as Single Ascending Doses in Healthy Subjects, and as a Single Dose in Healthy Subjects of Japanese-descent and in Subjects With Chronic Kidney Disease
This Phase I First in Human (FIH) study is being conducted to determine the safety, pharmacokinetics (PK), pharmacodynamics (PD), and immunogenicity profile of MEDI8367 across the dose range.
Start Date22 Jul 2020 |
Sponsor / Collaborator  AstraZeneca PLC AstraZeneca PLC [+1] |
100 Clinical Results associated with ITGB8
Login to view more data
100 Translational Medicine associated with ITGB8
Login to view more data
0 Patents (Medical) associated with ITGB8
Login to view more data
193
Literatures (Medical) associated with ITGB801 Dec 2024·Asia-Pacific Journal of Clinical Oncology
E74‐like factor 4 promotes the proliferation, invasion, and migration of colorectal cancer cells via long non‐coding RNA integrin subunit beta 8 antisense RNA 1‐mediated histone H3 lysine 27 trimethylation modification
Article
Author: Wang, Qi ; Liu, Shaofeng
Abstract:
Aim:
Colorectal cancer (CRC) is a common malignancy in the gastrointestinal tract. The main objective of this study is to explore the potential mechanisms of E74‐like factor 4 (ELF4) in CRC progression, providing a novel therapeutic target for CRC treatment.
Methods:
CRC cells and normal control cells were cultured. Levels of ELF4/long non‐coding RNA integrin subunit beta 8 antisense RNA 1 (LncRNA ITGB8‐AS1)/claudin‐23 (CLDN23) were detected by real‐time quantitative polymerase chain reaction or Western blot assay. ELF4 siRNA, ITGB8‐AS1 pcDNA3.1, or CLDN23 siRNA were transfected into cells. Cell proliferation, migration, and invasion were evaluated. The enrichment of ELF4 on the ITGB8‐AS1 promoter was detected. Dual‐luciferase assay was employed to assess the binding between ELF4 and the ITGB8‐AS1 promoter. RNA pull‐down and RNA immunoprecipitation assays were conducted to investigate the binding between ITGB8‐AS1 and enhancer of zeste homolog 2 (EZH2). The binding of EZH2 and histone H3 lysine 27 trimethylation (H3K27me3) to the CLDN23 promoter was detected.
Results:
ELF4 and ITGB8‐AS1 were upregulated in CRC cells, while CLDN23 was downregulated. Knockdown of ELF4 inhibited cell proliferation, invasion, and migration. Mechanistically, ELF4 transcriptionally activated ITGB8‐AS1 and promoted the binding between ITGB8‐AS1 and EZH2. EZH2 catalyzed H3K27me3 modification on the CLDN23 promoter, leading to decreased CLDN23 expression. Overexpression of ITGB8‐AS1 or downregulation of CLDN23 could reduce the inhibitory effects of silencing ELF4 on CRC cell proliferation, migration, and invasion.
Conclusion:
ELF4 accelerates CRC progression through the ITGB8‐AS1/CLDN23 axis, providing new therapeutic targets for CRC.
10 Sep 2024·Endocrine Metabolic & Immune Disorders-Drug Targets
Single-Cell RNA Sequencing Reveals Transcriptional Signatures and Cell-Cell Communication in Diabetic Retinopathy
Article
Author: Pang, Lin ; Peng, Yueling ; Li, Muye ; Wang, Lin ; Li, Junhong
Background::
Diabetic retinopathy (DR) is a major cause of vision loss in workingage
individuals worldwide. Cell-to-cell communication between retinal cells and retinal pigment
epithelial cells (RPEs) in DR is still unclear, so this study aimed to generate a single-cell atlas
and identify receptor‒ligand communication between retinal cells and RPEs.
Methods::
A mouse single-cell RNA sequencing (scRNA-seq) dataset was retrieved from the
GEO database (GSE178121) and was further analyzed with the R package Seurat. Cell cluster
annotation was performed to further analyze cell‒cell communication. The differentially expressed
genes (DEGs) in RPEs were explored through pathway enrichment analysis and the protein‒
protein interaction (PPI) network. Core genes in the PPI were verified by quantitative PCR
in ARPE-19 cells.
Results::
We observed an increased proportion of RPEs in STZ mice. Although some overall
intercellular communication pathways did not differ significantly in the STZ and control groups,
RPEs relayed significantly more signals in the STZ group. In addition, THBS1, ITGB1,
COL9A3, ITGB8, VTN, TIMP2, and FBN1 were found to be the core DEGs of the PPI network
in RPEs. qPCR results showed that the expression of ITGB1, COL9A3, ITGB8, VTN, TIMP2,
and FBN1 was higher and consistent with scRNA-seq results in ARPE-19 cells under hyperglycemic
conditions.
Conclusion::
Our study, for the first time, investigated how signals that RPEs relay to and from
other cells underly the progression of DR based on scRNA-seq. These signaling pathways and
hub genes may provide new insights into DR mechanisms and therapeutic targets.
01 Sep 2024·GENE
Preliminary study on the mechanism by which exosomes derived from human exfoliated deciduous teeth improve the proliferation and osteogenic inhibitory effect of glucocorticoid-induced BMSCs
Article
Author: Lv, Jian ; Xu, Jie ; Wang, Xinmin ; Bai, Yuxi ; Liu, Fei ; Lu, Yang
BACKGROUND:
Steroid-induced osteonecrosis of the femoral head (SONFH) is a disease characterized by a collapsed femoral head caused by the overuse of glucocorticoids. Dysfunction of bone marrow mesenchymal stem cells (BMSCs) is an important pathological feature of SONFH. In this study, we investigated whether exosomes from SHEDs (stem cells from human exfoliated deciduous teeth) have a therapeutic effect on glucocorticoid-induced inhibition of proliferation and osteogenesis in BMSCs, and elucidated the underlying mechanisms involved.
METHODS:
Primary dental pulp cells were isolated and cultured from human deciduous tooth pulp, SHEDs were isolated and purified by the limiting dilution method and exosomes were isolated from the supernatants of SHEDs by ultracentrifugation. The cell surface markers CD31, CD34, CD45, CD73, CD90 and CD105 were detected by flow cytometry. A Cell-Counting-Kit-8 assay was used to detect cell activity. ALP and Alizarin Red staining were used to identify osteogenic differentiation ability, and exosomes were identified using transmission electron microscopy, NanoFCM and Western blotting. PKH67 fluorescence was used to track the uptake of exosomes by BMSCs. Transcriptome analysis combined with quantitative real-time PCR was used to explore the underlying mechanism involved.
RESULTS:
Exosomes secreted by SHEDs can be endocytosed by BMSCs, and can partially reverse the inhibitory effects of glucocorticoids on the viability and osteogenic differentiation of BMSCs. Transcriptome sequencing analysis revealed that the differentially expressed mRNAs regulated by SHED-derived exosomes were enriched mainly in signaling pathways such as the apoptosis pathway, the PI3K-Akt signaling pathway, the Hippo signaling pathway and the p53 signaling pathway. qPCR showed that SHED-derived exosomes reversed the dexamethasone-induced upregulation of HGF and ITGB8 expression and the inhibition of EFNA1 expression, but further increased the dexamethasone-induced downregulation of IL7 expression. In conclusion, SHED-derived exosomes partially reversed the inhibitory effects of glucocorticoids on BMSC proliferation and osteogenesis by inhibiting the expression of HGF, ITGB8 and IL7, and upregulating the expression of EFNA1.
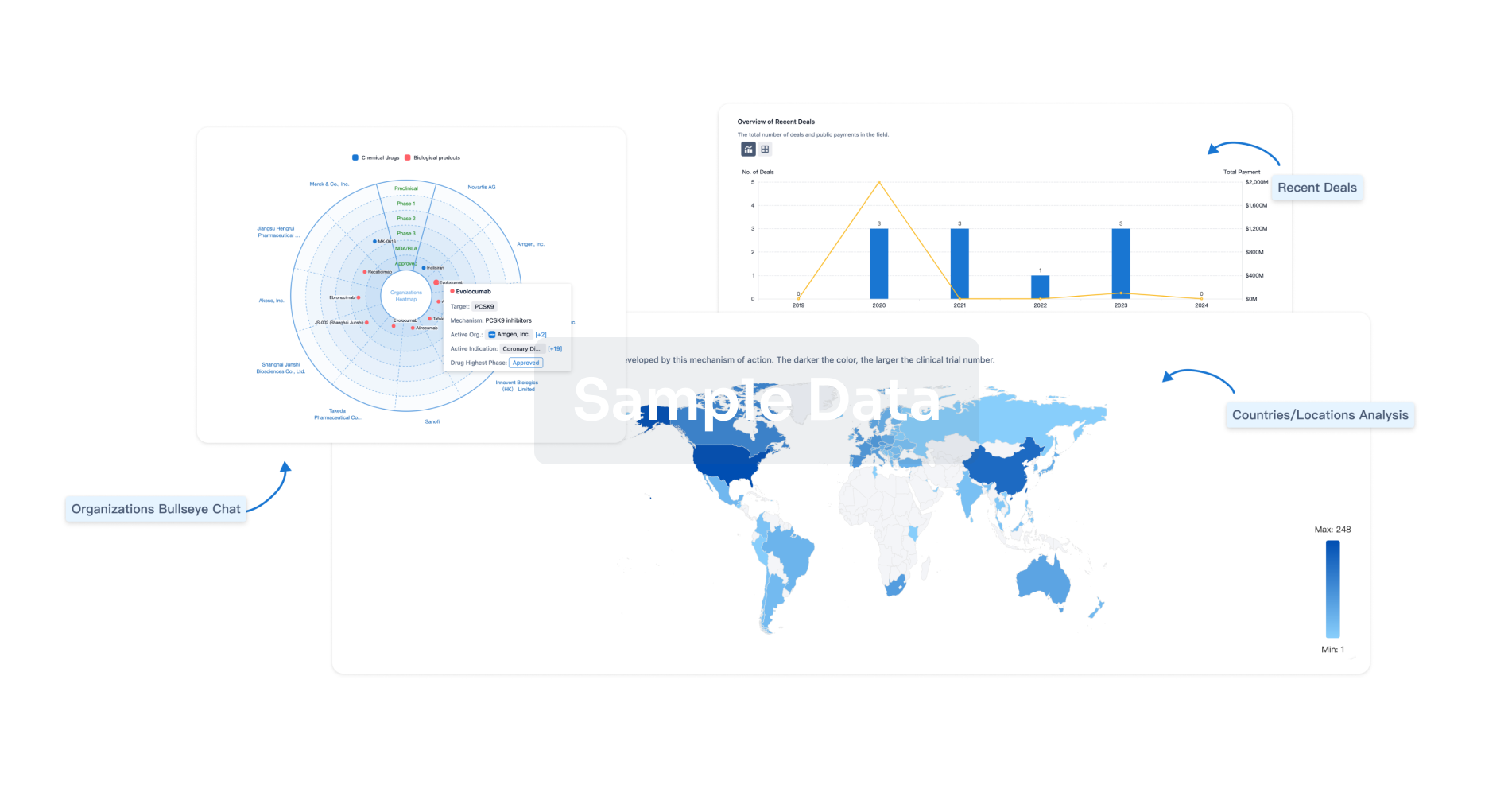
Analysis
Perform a panoramic analysis of this field.
login
or
AI Agents Built for Biopharma Breakthroughs
Accelerate discovery. Empower decisions. Transform outcomes.
Get started for free today!
Accelerate Strategic R&D decision making with Synapse, PatSnap’s AI-powered Connected Innovation Intelligence Platform Built for Life Sciences Professionals.
Start your data trial now!
Synapse data is also accessible to external entities via APIs or data packages. Empower better decisions with the latest in pharmaceutical intelligence.
Bio
Bio Sequences Search & Analysis
Sign up for free
Chemical
Chemical Structures Search & Analysis
Sign up for free


