Request Demo
Last update 08 May 2025
DIRAS1
Last update 08 May 2025
Basic Info
Synonyms Di-Ras1, DIRAS family GTPase 1, DIRAS1 + [6] |
Introduction Displays low GTPase activity and exists predominantly in the GTP-bound form. |
Related
100 Clinical Results associated with DIRAS1
Login to view more data
100 Translational Medicine associated with DIRAS1
Login to view more data
0 Patents (Medical) associated with DIRAS1
Login to view more data
28
Literatures (Medical) associated with DIRAS131 Dec 2024·Cancer Biology & Therapy
M6A modification regulates tumor suppressor DIRAS1 expression in cervical cancer cells
Article
Author: Wang, Yu-Yan ; Dai, Ning ; Zhang, Xin ; Gao, Wei-Ran ; Ye, Lian-Hua ; Yin, Yu ; Zhao, An-Qi
01 Dec 2024·Alzheimer's & Dementia
Identifying partial haplotypes that are associated with late‐onset Alzheimer disease
Article
Author: Climer, Sharlee
01 Jun 2023·The Journal of biological chemistry
GTPase splice variants RAC1 and RAC1B display isoform-specific differences in localization, prenylation, and interaction with the chaperone protein SmgGDS.
Article
Author: Williams, Carol L ; Auger, Shelby A ; Lorimer, Ellen ; Koehn, Olivia J ; Das, Akansha S ; Prokop, Jeremy W ; Harris, Ra'Mal ; Distefano, Mark D ; Unger, Bethany ; Suazo, Kiall F
Analysis
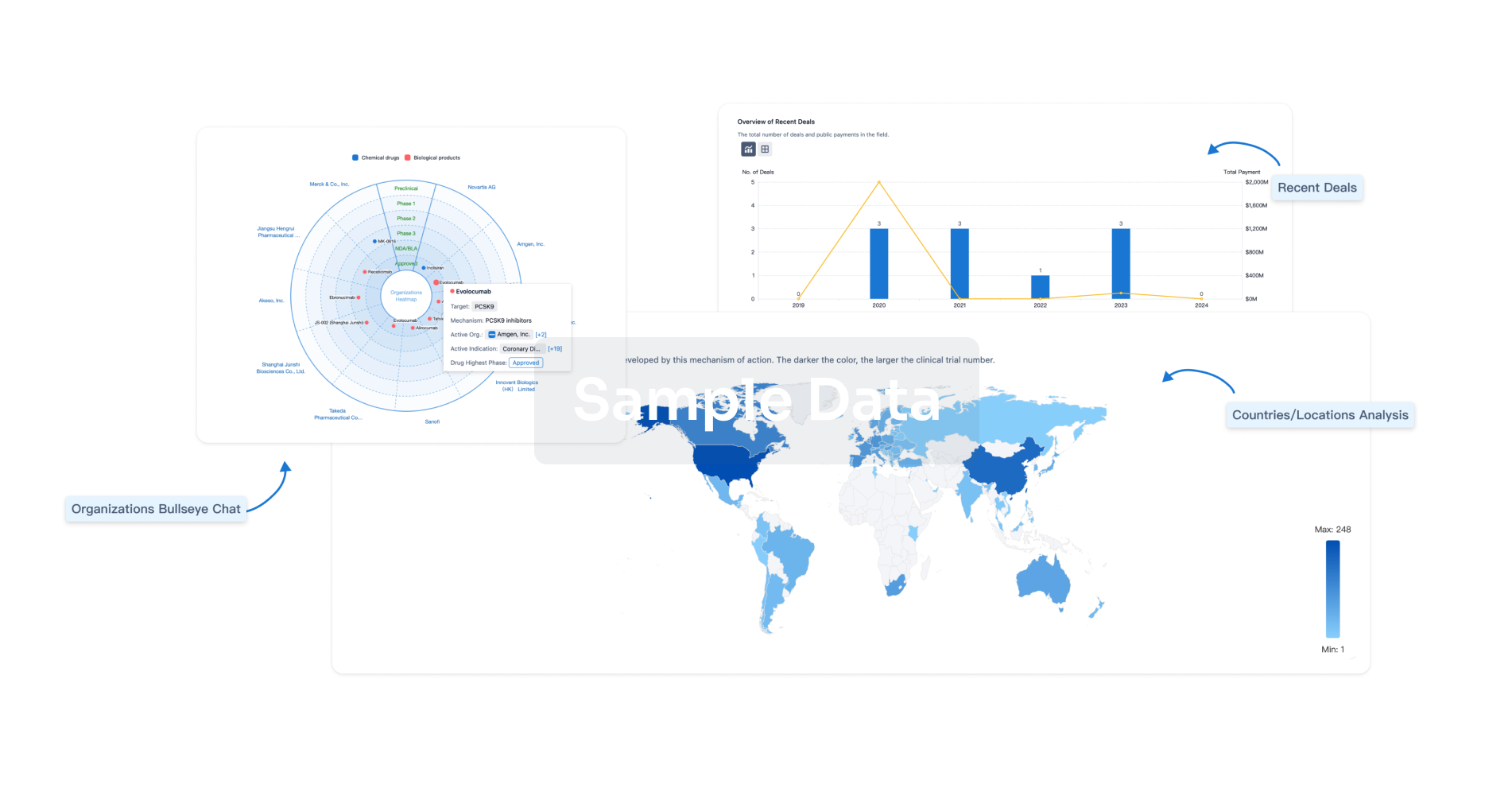
Perform a panoramic analysis of this field.
login
or
AI Agents Built for Biopharma Breakthroughs
Accelerate discovery. Empower decisions. Transform outcomes.
Get started for free today!
Accelerate Strategic R&D decision making with Synapse, PatSnap’s AI-powered Connected Innovation Intelligence Platform Built for Life Sciences Professionals.
Start your data trial now!
Synapse data is also accessible to external entities via APIs or data packages. Empower better decisions with the latest in pharmaceutical intelligence.
Bio
Bio Sequences Search & Analysis
Sign up for free
Chemical
Chemical Structures Search & Analysis
Sign up for free