Request Demo
Last update 08 May 2025
oxlT
Last update 08 May 2025
Basic Info
Synonyms- |
Introduction Anion transporter that carries out the exchange of divalent oxalate with monovalent formate, the product of oxalate decarboxylation, at the plasma membrane, and in doing so catalyzes the vectorial portion of a proton-motive metabolic cycle that drives ATP synthesis. |
Related
1
Drugs associated with oxlTMechanism frc stimulants [+5] |
Active Org.- |
Originator Org. |
Active Indication- |
Inactive Indication |
Drug Highest PhasePending |
First Approval Ctry. / Loc.- |
First Approval Date20 Jan 1800 |
2
Clinical Trials associated with oxlTNCT05377112
A Double-Blind, Randomized, Placebo-controlled Study to Assess the Safety, Tolerability, and Pharmacodynamics of SYNB8802v1 in Subjects With History of Gastric Bypass Surgery or Short-bowel Syndrome
Study SYNB8802-CP-002 is designed to assess safety, tolerability, and oxalate lowering, in subjects with a history of gastric bypass surgery or short-bowel syndrome. In addition, this study will explore other PD effects relative to baseline as well as predictors of efficacy and tolerability.
Start Date29 Mar 2022 |
Sponsor / Collaborator |
NCT04629170
A Double-Blind, Randomized, Placebo-Controlled Study to Assess the Safety, Tolerability, and Pharmacodynamics of SYNB8802 in Healthy Volunteers and in Patients With Enteric Hyperoxaluria
This Phase 1a/b, first-in-human, multiple dose-escalation, randomized, double-blinded, placebo-controlled study is evaluating SYNB8802 in healthy volunteers (HV) and subjects diagnosed with enteric hyperoxaluria (EH). Eligible subjects receive investigational product (IP) and undergo safety monitoring, evaluations, and subsequent follow-up after IP administration.
In Part 2, all evaluations and assessments throughout this study may be conducted either at the clinical site or by a home healthcare professional at an alternative location (e.g., EH patient's home, hotel).
In Part 2, all evaluations and assessments throughout this study may be conducted either at the clinical site or by a home healthcare professional at an alternative location (e.g., EH patient's home, hotel).
Start Date04 Nov 2020 |
Sponsor / Collaborator |
100 Clinical Results associated with oxlT
Login to view more data
100 Translational Medicine associated with oxlT
Login to view more data
0 Patents (Medical) associated with oxlT
Login to view more data
32
Literatures (Medical) associated with oxlT25 Jan 2024·The Journal of Physical Chemistry Letters
Accelerated Molecular Dynamics and AlphaFold Uncover a Missing Conformational State of Transporter Protein OxlT
Article
Author: Jaunet-Lahary, Titouan ; Okazaki, Kei-Ichi ; Yamashita, Atsuko ; Ohnuki, Jun
01 Jan 2023·Frontiers in microbiology
Expanding the taxonomic and environmental extent of an underexplored carbon metabolism-oxalotrophy.
Article
Author: Sonke, Alexander ; Trembath-Reichert, Elizabeth
01 Oct 2021·Protein ScienceQ4 · BIOLOGY
An optogenetic assay method for electrogenic transporters using Escherichia coli co‐expressing light‐driven proton pump
Q4 · BIOLOGY
Article
Author: Yamashita, Atsuko ; Hayashi, Masahiro ; Kojima, Keiichi ; Sudo, Yuki
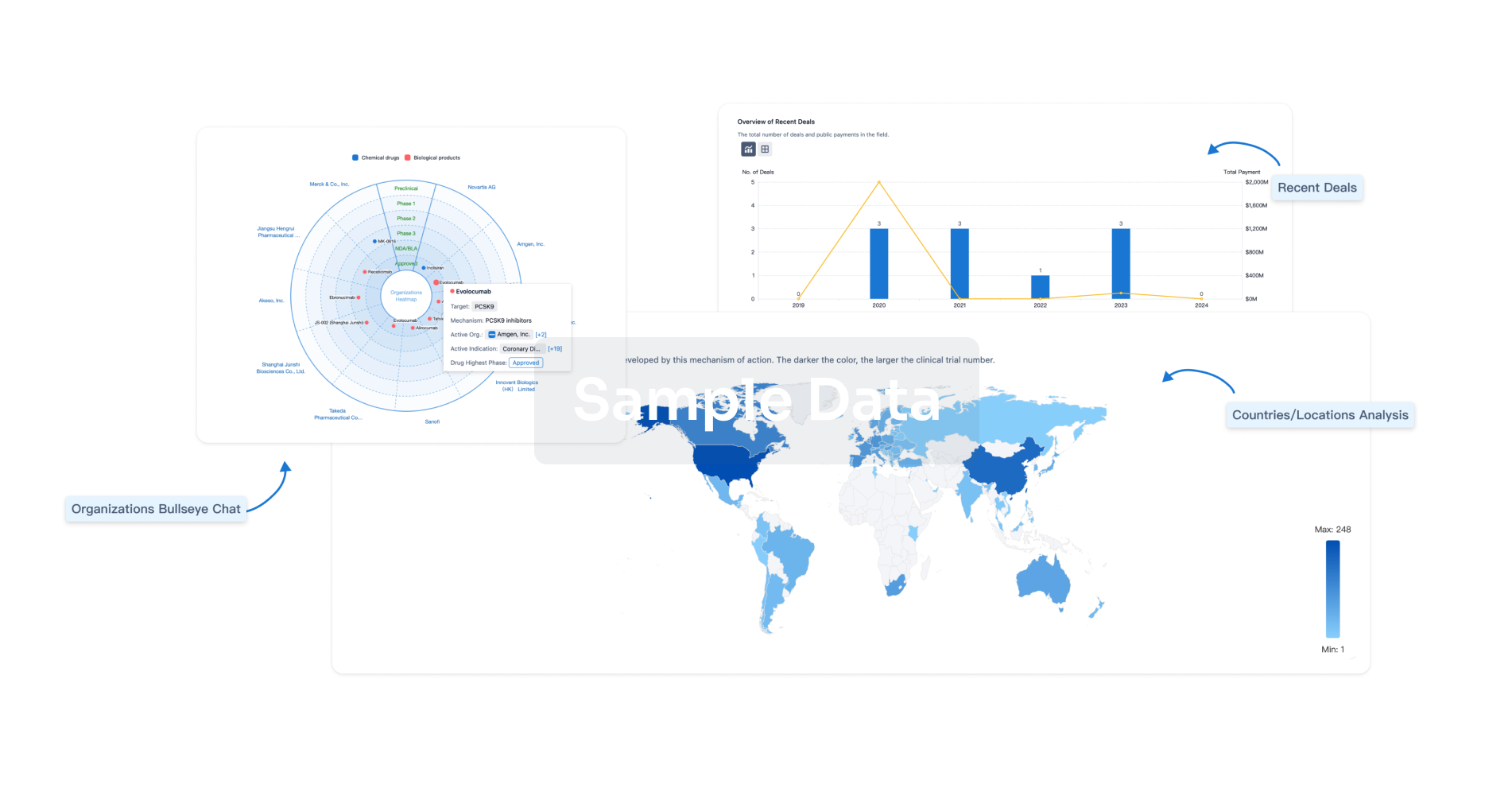
Analysis
Perform a panoramic analysis of this field.
login
or
AI Agents Built for Biopharma Breakthroughs
Accelerate discovery. Empower decisions. Transform outcomes.
Get started for free today!
Accelerate Strategic R&D decision making with Synapse, PatSnap’s AI-powered Connected Innovation Intelligence Platform Built for Life Sciences Professionals.
Start your data trial now!
Synapse data is also accessible to external entities via APIs or data packages. Empower better decisions with the latest in pharmaceutical intelligence.
Bio
Bio Sequences Search & Analysis
Sign up for free
Chemical
Chemical Structures Search & Analysis
Sign up for free
