Request Demo
Last update 08 May 2025
NOC2L
Last update 08 May 2025
Basic Info
Synonyms DKFZP564C186, NET15, NET7 + [8] |
Introduction Acts as an inhibitor of histone acetyltransferase activity; prevents acetylation of all core histones by the EP300/p300 histone acetyltransferase at p53/TP53-regulated target promoters in a histone deacetylases (HDAC)-independent manner. Acts as a transcription corepressor of p53/TP53- and TP63-mediated transactivation of the p21/CDKN1A promoter. Involved in the regulation of p53/TP53-dependent apoptosis. Associates together with TP63 isoform TA*-gamma to the p21/CDKN1A promoter. |
Analysis
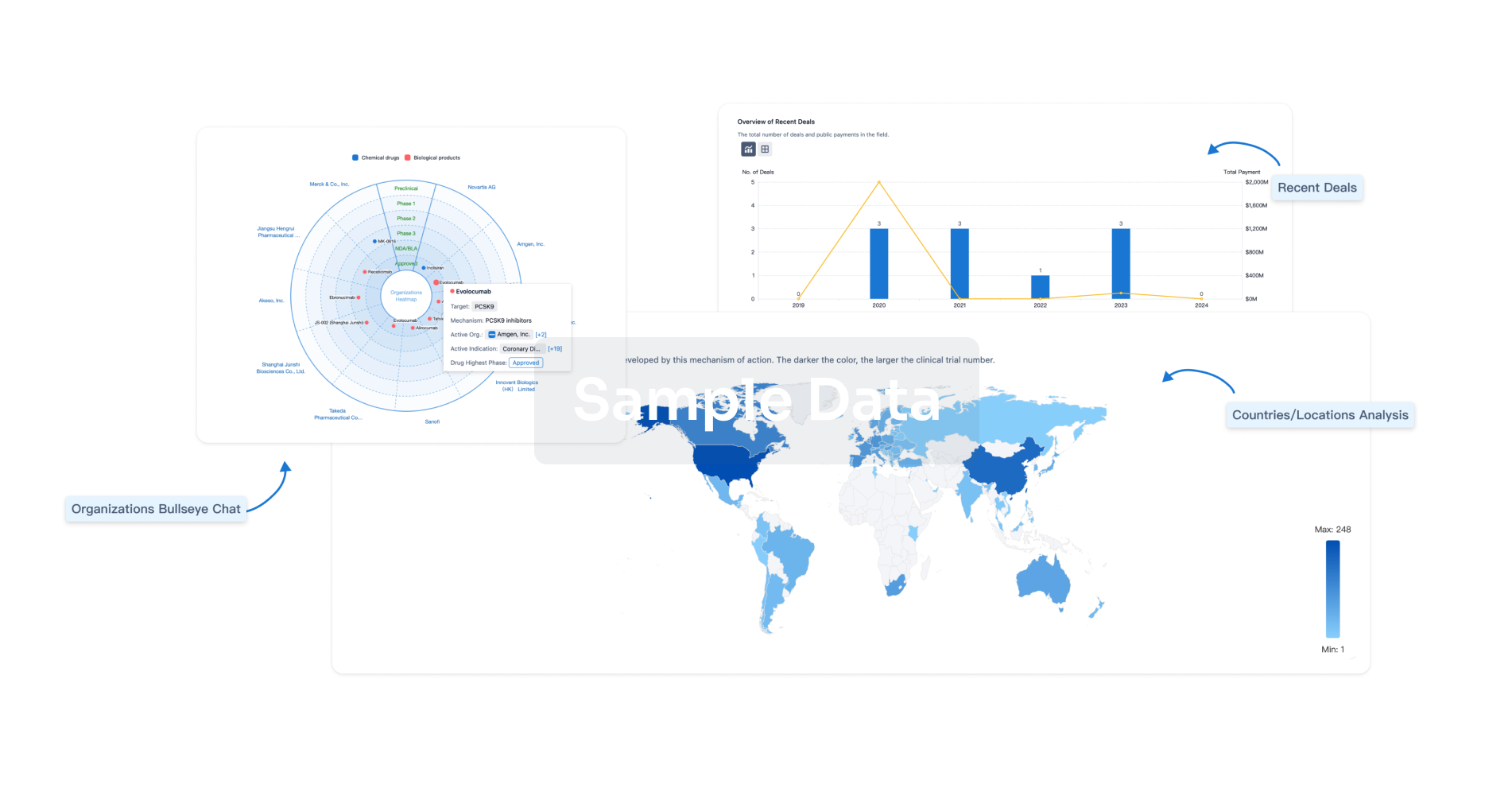
Perform a panoramic analysis of this field.
login
or
AI Agents Built for Biopharma Breakthroughs
Accelerate discovery. Empower decisions. Transform outcomes.
Get started for free today!
Accelerate Strategic R&D decision making with Synapse, PatSnap’s AI-powered Connected Innovation Intelligence Platform Built for Life Sciences Professionals.
Start your data trial now!
Synapse data is also accessible to external entities via APIs or data packages. Empower better decisions with the latest in pharmaceutical intelligence.
Bio
Bio Sequences Search & Analysis
Sign up for free
Chemical
Chemical Structures Search & Analysis
Sign up for free