Request Demo
Last update 08 May 2025
DOCK3
Last update 08 May 2025
Basic Info
Synonyms dedicator of cytokinesis 3, Dedicator of cytokinesis protein 3, DOCK3 + [5] |
Introduction Potential guanine nucleotide exchange factor (GEF). GEF proteins activate some small GTPases by exchanging bound GDP for free GTP. Its interaction with presenilin proteins as well as its ability to stimulate Tau/MAPT phosphorylation suggest that it may be involved in Alzheimer disease. Ectopic expression in nerve cells decreases the secretion of amyloid-beta APBA1 protein and lowers the rate of cell-substratum adhesion, suggesting that it may affect the function of some small GTPase involved in the regulation of actin cytoskeleton or cell adhesion receptors (By similarity). |
Analysis
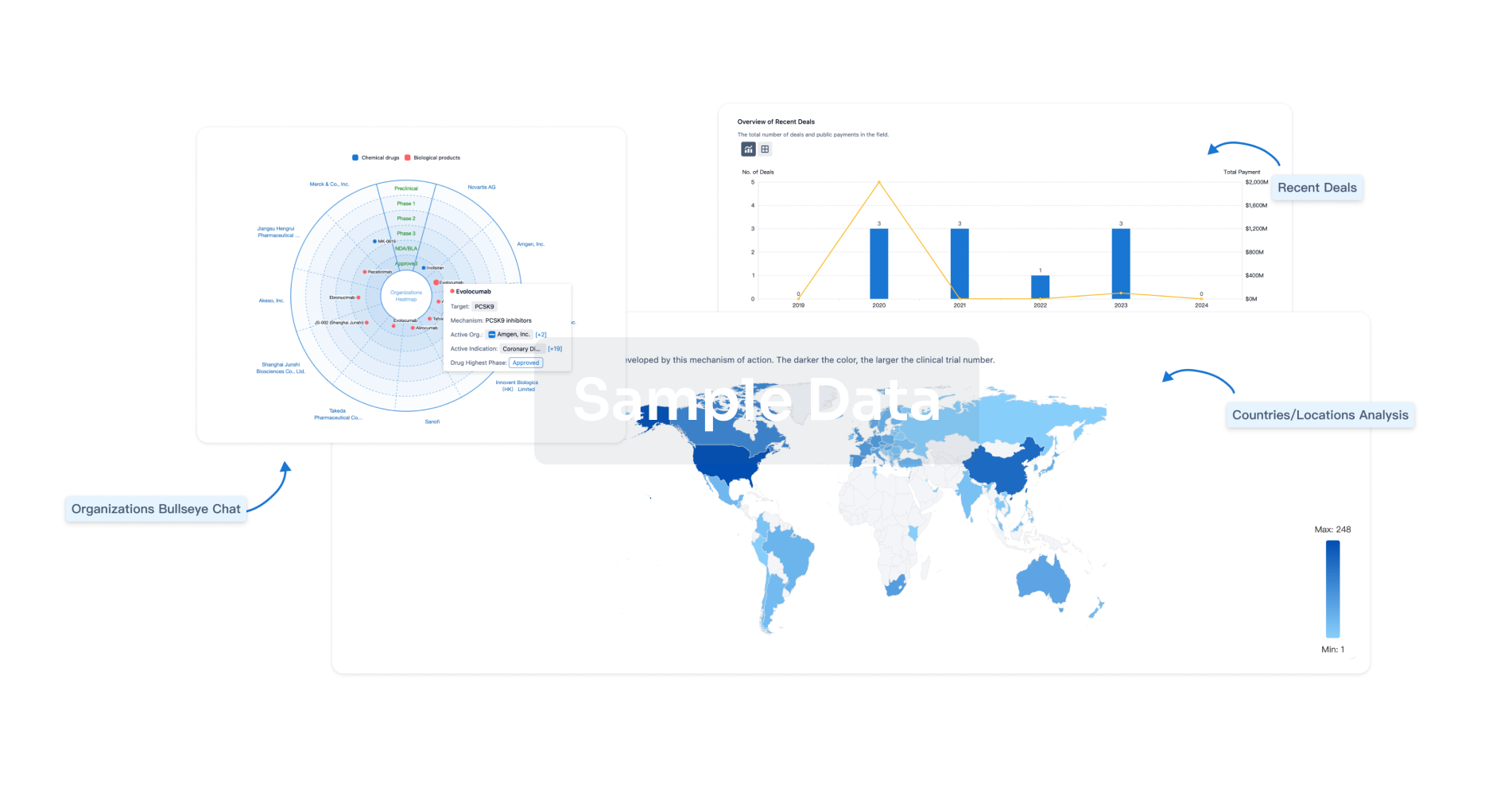
Perform a panoramic analysis of this field.
login
or
AI Agents Built for Biopharma Breakthroughs
Accelerate discovery. Empower decisions. Transform outcomes.
Get started for free today!
Accelerate Strategic R&D decision making with Synapse, PatSnap’s AI-powered Connected Innovation Intelligence Platform Built for Life Sciences Professionals.
Start your data trial now!
Synapse data is also accessible to external entities via APIs or data packages. Empower better decisions with the latest in pharmaceutical intelligence.
Bio
Bio Sequences Search & Analysis
Sign up for free
Chemical
Chemical Structures Search & Analysis
Sign up for free