Request Demo
Last update 08 May 2025
SRC x Raf kinase
Last update 08 May 2025
Basic Info
Related Targets |
Related
7
Drugs associated with SRC x Raf kinaseTarget |
Mechanism Raf kinase inhibitors [+1] |
Originator Org. |
Active Indication |
Inactive Indication |
Drug Highest PhasePhase 1 |
First Approval Ctry. / Loc.- |
First Approval Date20 Jan 1800 |
Target |
Mechanism Raf kinase inhibitors [+1] |
Active Org. |
Originator Org. |
Active Indication |
Inactive Indication- |
Drug Highest PhasePreclinical |
First Approval Ctry. / Loc.- |
First Approval Date20 Jan 1800 |
Mechanism BRAF inhibitors [+3] |
Active Org. |
Originator Org. |
Active Indication |
Inactive Indication- |
Drug Highest PhasePreclinical |
First Approval Ctry. / Loc.- |
First Approval Date20 Jan 1800 |
1
Clinical Trials associated with SRC x Raf kinaseNCT02437227
A Phase 1, First in Man, Dual Centre, Open-label Dose Escalation Study With Expansion to Evaluate the Safety, Tolerability, Pharmacokinetics and Pharmacodynamics of CCT3833 (BAL3833), a panRAF Inhibitor, Given Orally in Patients With Advanced Solid Tumours, Including Metastatic Melanoma
The study is a first in man, dose escalation study to evaluate the safety, tolerability and how the drug works in the body in patients with all solid tumours. The aim of this study is to determine the most effective dose of the study drug that can then be further investigated in patients with advanced melanoma.
Start Date15 Apr 2015 |
Sponsor / Collaborator |
100 Clinical Results associated with SRC x Raf kinase
Login to view more data
100 Translational Medicine associated with SRC x Raf kinase
Login to view more data
0 Patents (Medical) associated with SRC x Raf kinase
Login to view more data
546
Literatures (Medical) associated with SRC x Raf kinase01 Jul 2025·Brain Research
Effects of Asiatic acid on brain cancer by altering astrocytes and the AKT1-PRKCB signaling pathway: A genomic and network pharmacology perspective
Article
Author: Kumar Mishra, Sunil ; Singh, Amit Kumar ; Agrawal, Shreni ; Tiwari, Kavindra Nath ; Kumar, Pradeep ; Kumar Pathak, Adarsh ; Kumar Sah Gond, Manjeet ; Kumar Singh, Anand ; Singh Negi, Rohit ; Das, Richa
01 May 2025·Toxicology
A network toxicology and molecular docking-based approach revealed shared hepatotoxic mechanisms and targets between the herbicide 2,4-D and its metabolite 2,4-DCP
Article
Author: Gomes, Cleyton ; Farias, Davi ; Martins, Rafael Xavier ; Souza, Juliana Alves da Costa Ribeiro ; Carvalho, Matheus ; Souza, Terezinha
08 Nov 2024·Medicine
Network pharmacology and molecular docking to explore the potential molecular mechanism of chlorogenic acid treatment of oral squamous cell carcinoma
Article
Author: Hao, Puyu ; Xve, Xulong ; Zhang, Jun ; Yang, Yutao ; Feng, Zhanqin
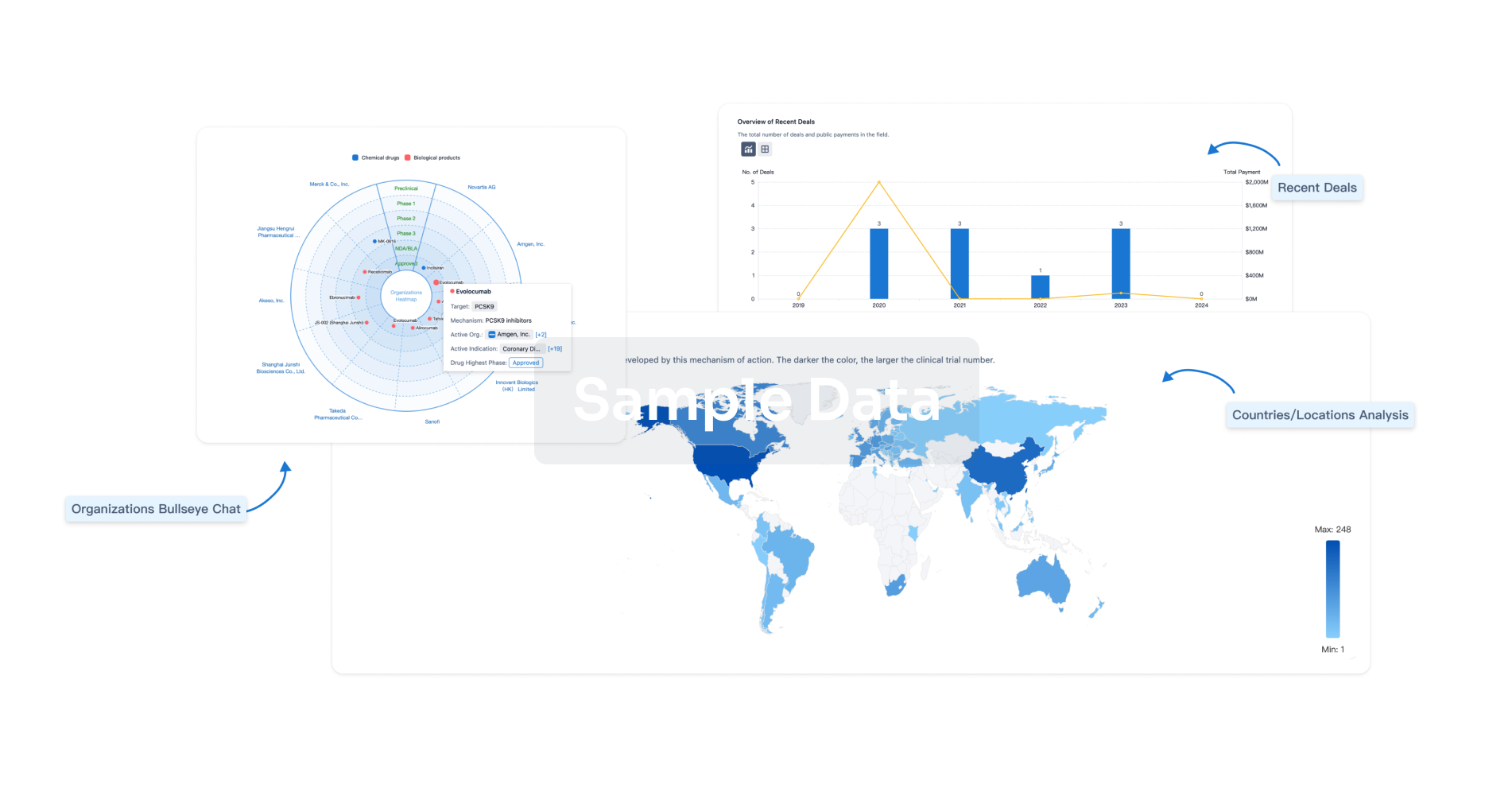
Analysis
Perform a panoramic analysis of this field.
login
or
AI Agents Built for Biopharma Breakthroughs
Accelerate discovery. Empower decisions. Transform outcomes.
Get started for free today!
Accelerate Strategic R&D decision making with Synapse, PatSnap’s AI-powered Connected Innovation Intelligence Platform Built for Life Sciences Professionals.
Start your data trial now!
Synapse data is also accessible to external entities via APIs or data packages. Empower better decisions with the latest in pharmaceutical intelligence.
Bio
Bio Sequences Search & Analysis
Sign up for free
Chemical
Chemical Structures Search & Analysis
Sign up for free




