Request Demo
Last update 09 Sep 2025

OPKO Health, Inc.
Last update 09 Sep 2025
Overview
Tags
Endocrinology and Metabolic Disease
Other Diseases
Nervous System Diseases
Hormone
CTP fusion protein
Synthetic peptide
Disease domain score
A glimpse into the focused therapeutic areas
No Data
Technology Platform
Most used technologies in drug development
No Data
Targets
Most frequently developed targets
No Data
| Top 5 Drug Type | Count |
|---|---|
| Small molecule drug | 5 |
| Synthetic peptide | 4 |
| Multi-specific antibody | 4 |
| ASO | 1 |
| Gene therapy | 1 |
Related
23
Drugs associated with OPKO Health, Inc.Target |
Mechanism GHR agonists |
Active Org. |
Originator Org. |
Inactive Indication- |
Drug Highest PhaseApproved |
First Approval Ctry. / Loc. Canada |
First Approval Date26 Oct 2021 |
Target |
Mechanism VDR agonists |
Active Org. |
Originator Org. |
Active Indication |
Inactive Indication |
Drug Highest PhaseApproved |
First Approval Ctry. / Loc. United States |
First Approval Date05 Aug 1980 |
Target |
Mechanism APP inhibitors |
Active Org. |
Originator Org. |
Active Indication |
Inactive Indication |
Drug Highest PhasePhase 3 |
First Approval Ctry. / Loc.- |
First Approval Date- |
55
Clinical Trials associated with OPKO Health, Inc.NCT07110584
A Phase 1/2, Multi-Center, Open-Label Clinical Study Evaluating MDX2004 In Participants With Advanced Tumors
This study is designed to characterize the safety, tolerability, and anti-tumor activity of MDX2004 in patients with advanced tumors.
Start Date30 Sep 2025 |
Sponsor / Collaborator |
NCT06239194
A Phase 1/2a, Multicenter, First-in-human, Open-label Clinical Trial Evaluating MDX2001 Monotherapy in Patients With Advanced Solid Tumors
This study is designed to characterize the safety, tolerability, and anti-tumor activity of MDX2001 in patients with advanced solid tumors.
Start Date12 Jun 2024 |
Sponsor / Collaborator |
CTR20221404
一项评估IDX-1197联合XeIox(卡培他滨和奥沙利铂)或伊立替康用于晚期胃癌患者的安全性、耐受性和疗效的开放性、国际、多中心、Ib期研究
[Translation] An open-label, international, multicenter, phase Ib study evaluating the safety, tolerability, and efficacy of IDX-1197 in combination with XeIox (capecitabine and oxaliplatin) or irinotecan in patients with advanced gastric cancer
研究的主要目的是评价IDX-1197的安全性和耐受性,并确定联合卡培他滨和奥沙利铂(Xelox),或伊立替康用于治疗晚期胃癌患者的最大耐受剂量(MTD)和推荐II期剂量(RP2D)。 研究的次要目的为:评估IDX-1197与Xelox或伊立替康联合给药时的疗效,疗效是通过按实体瘤疗效评价标准(RECIST)v1.1评价的客观缓解率(ORR)、无进展生存期(PFS)、疾病控制率(DCR)、缓解持续时间(DOR)、缓解时间(TTR)、最佳总缓解(BOR),疾病进展时间(TTP),最低可重复有效剂量(MRAD)以及总生存期(OS)进行定义。评价IDX-1197单次递增剂量(SAD)联合Xelox或伊立替康用于治疗晚期胃癌患者时的药代动力学(PK)(仅剂量探索部分)。评价IDX-1197联合Xelox或伊立替康用于治疗晚期胃癌患者时IDX-1197和伊立替康的群体PK模型。通过评估校正的QT间期,评价IDX-1197联合伊立替康对心脏复极化的影响。通过ECG记录和时间匹配的PK测量来评估QT间期延长的风险。
[Translation]
The primary objectives of the study are to evaluate the safety and tolerability of IDX-1197 and to determine the maximum tolerated dose (MTD) and recommended phase II dose (RP2D) of IDX-1197 in combination with capecitabine and oxaliplatin (Xelox) or irinotecan for the treatment of patients with advanced gastric cancer. Secondary objectives of the study are to evaluate the efficacy of IDX-1197 in combination with Xelox or irinotecan, as defined by objective response rate (ORR), progression-free survival (PFS), disease control rate (DCR), duration of response (DOR), time to response (TTR), best overall response (BOR), time to disease progression (TTP), lowest repeatable effective dose (MRAD), and overall survival (OS) as assessed by response evaluation criteria in solid tumors (RECIST) v1.1. To evaluate the pharmacokinetics (PK) of IDX-1197 in combination with Xelox or irinotecan as a single ascending dose (SAD) in patients with advanced gastric cancer (dose-finding portion only). Population PK models of IDX-1197 and irinotecan were evaluated in combination with Xelox or irinotecan in patients with advanced gastric cancer. The effects of IDX-1197 plus irinotecan on cardiac repolarization were evaluated by assessing the corrected QT interval. The risk of QT prolongation was assessed by ECG recordings and time-matched PK measurements.
Start Date15 Dec 2023 |
Sponsor / Collaborator  Idience Co., Ltd. Idience Co., Ltd. [+2] |
100 Clinical Results associated with OPKO Health, Inc.
Login to view more data
0 Patents (Medical) associated with OPKO Health, Inc.
Login to view more data
52
Literatures (Medical) associated with OPKO Health, Inc.09 Oct 2024·Science Translational Medicine
A multispecific antibody against SARS-CoV-2 prevents immune escape in vitro and confers prophylactic protection in vivo
Article
Author: Kwong, Peter D. ; Zhang, Yi ; Oloniniyi, Olamide K. ; Boruszczak, Marika ; Andersen, Hanne ; Yang, Zhi-Yong ; Douek, Daniel C. ; Harris, Darcy R. ; Pegu, Amarendra ; Wei, Ronnie R. ; Wang, Lingshu ; Choe, Misook ; Porto, Maciel ; Godbole, Sucheta ; Shi, Wei ; Wei, Chih-Jen ; Zhao, Bingchun ; Moliva, Juan I. ; Wolff, Jeremy J. ; Suthar, Mehul S. ; Nabel, Gary J. ; Leung, Kwanyee ; Henry, Amy R. ; Zhou, Tongqing ; Olia, Adam S. ; Longobardi, Lindsay ; Carlton, Kevin ; Yang, Eun Sung ; Li, Juan ; Sullivan, Nancy J. ; Misasi, John ; Chen, Man ; Pessaint, Laurent ; Nase, Danielle ; Mascola, John R. ; Gall, Jason G. ; Lei, Q. Paula ; Ivleva, Vera B. ; Merriam, Jonah S. ; Koup, Richard A. ; Wallace, Shannon M. ; Liu, Cuiping
Despite effective countermeasures, severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) persists worldwide because of its ability to diversify and evade human immunity. This evasion stems from amino acid substitutions, particularly in the receptor binding domain (RBD) of the spike protein that confers resistance to vaccine-induced antibodies and antibody therapeutics. To constrain viral escape through resistance mutations, we combined antibody variable regions that recognize different RBD sites into multispecific antibodies. Here, we describe multispecific antibodies, including a trivalent trispecific antibody that potently neutralized diverse SARS-CoV-2 variants and prevented virus escape more effectively than single antibodies or mixtures of the parental antibodies. Despite being generated before the appearance of Omicron, this trispecific antibody neutralized all major Omicron variants through BA.4/BA.5 at nanomolar concentrations. Negative stain electron microscopy suggested that synergistic neutralization was achieved by engaging different epitopes in specific orientations that facilitated binding across more than one spike protein. Moreover, a tetravalent trispecific antibody containing the same variable regions as the trivalent trispecific antibody also protected Syrian hamsters against Omicron variants BA.1, BA.2, and BA.5 challenge, each of which uses different amino acid substitutions to mediate escape from therapeutic antibodies. These results demonstrated that multispecific antibodies have the potential to provide broad SARS-CoV-2 coverage, decrease the likelihood of escape, simplify treatment, and provide a strategy for antibody therapies that could help eliminate pandemic spread for this and other pathogens.
01 Apr 2024·Environmental research
Metal oxide nanomaterials based electrochemical and optical biosensors for biomedical applications: Recent advances and future prospectives
Review
Author: Kumar, Parveen ; Liu, Bo ; Huo, Peipei ; Behl, Gautam ; Upadhyaya, Kapil ; Rajan, Ramachandran ; Xiang, Xin-Xin
The amalgamation of nanostructures with modern electrochemical and optical techniques gave rise to interesting devices, so-called biosensors. A biosensor is an analytical tool that incorporates various biomolecules with an appropriate physicochemical transducer. Over the past few years, metal oxide nanomaterials (MONMs) have significantly stimulated biosensing research due to their desired functionalities, versatile chemical stability, and low cost along with their unique optical, catalytic, electrical, and adsorption properties that provide an attractive platform for linking the biomolecules, for example, antibodies, nucleic acids, enzymes, and receptor proteins as sensing elements with the transducer for the detection of signals or signal amplifications. The signals to be measured are in direct proportionate to the concentration of the bioanalyte. Because of their simplicity, cost-effectiveness, portability, quick analysis, higher sensitivity, and selectivity against a broad range of biosamples, MONMs-based electrochemical and optical biosensing platforms are exhaustively explored as powerful early-diagnosis tools for point of care applications. Herein, we made a bibliometric analysis of past twenty years (2004-2023) on the application of MONMs as electrochemical and optical biosensing units using Web of Science database and the results of which clearly reveal the increasing number of publications since 2004. Geographical area distribution analysis of these publications shows that China tops the list followed by the United States of America and India. In this review, we first describe the electrochemical and optical properties of MONMs that are crucial for the creation of extremely stable, specific, and sensitive sensors with desirable characteristics. Then, the biomedical applications of MONMs-based bare and hybrid electrochemical and optical biosensing frameworks are highlighted in the light of recent literature. Finally, current limitations and future challenges in the field of biosensing technology are addressed.
01 Apr 2024·International journal of pharmaceutics
Design of liposomal nanocarriers with a potential for combined dexamethasone and bevacizumab delivery to the eye
Article
Author: Gasco, Paolo ; Fitzhenry, Laurence ; Behl, Gautam ; O'Reilly, Niall J ; Raghu Raj Singh, Thakur ; Ahmed, Zubair ; Farooq, Umer ; McLoughlin, Peter
Clinicians face numerous challenges when delivering medications to the eyes topically because of physiological barriers, that can inhibit the complete dose from getting to the intended location. Due to their small size, the ability to deliver drugs of different polarities simultaneously, and their biocompatibility, liposomes hold great promise for ocular drug delivery. This study aimed to develop and characterise a dual loaded liposome formulation encapsulating Bevacizumab (BEV) and Dexamethasone (DEX) that possessed the physicochemical attributes suitable for topical ocular delivery. Liposomes were prepared by using thin film hydration followed by extrusion, and the formulations were optimised using a design of experiments approach. Physicochemical characterisation along with cytocompatibility and bioactivity of the formulations were assessed. Liposomes were successfully prepared with a particle size of 139 ± 2 nm, PDI 0.03 ± 0.01 and zeta potential -2 ± 0.7 mV for the optimised formulation. BEV and DEX were successfully encapsulated into the liposomes with an encapsulation efficiency of 97 ± 0.5 % and 26 ± 0.5 %, respectively. A sustained release of BEV was observed from the liposomes and the bioactivity of the formulation was confirmed using a wound healing assay. In summary, a potential topical eye drop drug delivery system, which can co-load DEX and BEV was developed and characterised for its potential to be used in ocular drug delivery.
192
News (Medical) associated with OPKO Health, Inc.14 Aug 2025
Company Reports $13.2M in Q2 Revenue, Achieves Significant Reduction in Operating Expenses and Continues Cost Reduction and Efficiency Improvements to Support Growth
COCONUT GROVE, Fla., Aug. 14, 2025 /PRNewswire/ -- NextPlat Corp (NASDAQ: NXPL, NXPLW) ("NextPlat" or the "Company"), a global consumer products and services company providing healthcare and technology solutions through eCommerce and retail channels worldwide, today announced the financial results for the quarter-ended June 30, 2025, reflecting the performance of its e-Commerce and Healthcare Operations.
"Results for the second quarter reflect continued top-line trends in the business, highlighted by strong e-Commerce growth and challenges in our Healthcare Operations as we continue to implement activities across the business to improve our operations, benefits that we believe are just beginning to be demonstrated by a reduced cost structure," said David Phipps, Interim CEO, President and CEO of Global Operations at NextPlat Corp. "Since the close of the second quarter, NextPlat continued to make progress against its plans to maximize the value of its business and adapt to the domestic and international challenges which have impacted us over the past few months. Our teams have been working hard to implement a broad array of initiatives within our Healthcare Operations which we believe have the potential to positively transform our business while enabling us to better attract and retain our customers. We are especially excited about the business expansion opportunities we see in the near-term and the new relationships we are working to form which will support our efforts to move into higher margin and higher growth segments of the Healthcare spectrum," said Mr. Phipps.
Second Quarter 2025 Financial Highlights:
Consolidated revenue for the quarter ended June 30, 2025, was approximately $13.2 million compared to approximately $17.0 million for the quarter ended June 30, 2024. The decrease in consolidated revenue was primarily driven by a decline in Healthcare Operations due to the following:
A $2.3 million decrease in pharmacy prescription and other revenue, net of PBM fees, to approximately $8.2 million for the quarter ended June 30, 2025 when compared to approximately $10.5 million in the prior year quarter, resulted from the decrease in pharmacy prescriptions filled which was influenced in part by the continued changes in provider relationships and shifts in patient flow due to insurance network adjustments or provider decisions to align with different pharmacy partners.
A $2.0 million decrease in pharmacy 340B contract revenue to approximately $1.0 million for the quarter ended June 30, 2025 when compared to approximately $3.0 million in the prior year quarter, resulted from certain relationships transitioning to other pharmacy partners, some covered entities opened in-house pharmacies, and another covered entity no longer participates in the 340B program.
Overall gross profit margin for the quarter ended June 30, 2025, declined to approximately 21.8% from 34.5% when compared to the prior year quarter. Gross profit margin from our Healthcare segment decreased to approximately 19.9% in the second quarter of 2025 from 35.2% in the second quarter of 2024 and was primarily attributable to the decrease in pharmacy prescriptions filled and 340B revenue as well as the continued industry-wide impact of drug price increases outpacing reimbursement rate adjustments. Gross profit margin for e-Commerce Operations decreased to approximately 25.9% from 31.6% when compared to the prior year quarter primarily due to a service provider airtime contract that expired on December 31, 2024, which introduced new airtime costs beginning January 1, 2025, and temporary rate reductions for some customers continuing to be affected by ongoing network service interruptions.
Operating expenses for the quarter ended June 30, 2025, decreased to approximately $4.7 million compared to approximately $16.8 million in the prior year quarter which included approximately $9.7 million in non-recurring expenses such as an impairment loss from the write-down of certain long-lived assets. As expected, overall operational costs decreased due to a decrease in stock-based compensation for grants fully vested a reduction in executive compensation, and ongoing headcount reductions. The Company has undertaken a series of proactive steps designed to improve its expense structure. As a result of ongoing operational process efficiency improvements, staff adjustments and other cost reduction efforts, the Company anticipates annual expense savings in excess of approximately $1.0 million, a portion of which management intends to reinvest into the business to support the launch of new and expanded service offerings and drive revenue growth.
Net loss attributable to NextPlat Corp common shareholders for the quarter ended June 30, 2025, decreased 66% to approximately $1.8 million, or ($0.07) per diluted share, compared to a net loss of approximately $5.3 million, or ($0.28) diluted earnings per share reported for the quarter ending June 30, 2024.
The Company ended the quarter with approximately $16.6 million in cash.
Organizational Highlights and Recent Business Developments:
During the second quarter, the Company continued its ongoing efforts to implement cost-reduction and business process improvements in its Healthcare Operations such as technology upgrades, and the recruitment of dedicated sales professionals focused on opportunities in the 340B and Long-Term Care segment. The positive impact of these efforts is expected to contribute to operational results throughout the remainder of 2025 and into 2026.
Following the passing of our former CEO and Chairman, Charles M. Fernandez, and as outlined in the Interim CEO Shareholder Update Letter dated June 30, 2025, the Company completed an extensive review of the business with a particular focus on its Healthcare Operations, which currently represents the largest component of revenue. As a result, the Company has identified three primary objectives: continued operational process efficiency and cost reduction efforts to drive maximum ROI; ensuring that the necessary talent is leading the business and that they are empowered to manage and drive growth; and, renewing a commitment to prudently invest in the business to help capitalize on opportunities, both organic and non-organic, while enhancing cashflow and long-term profitability. The Company expects to announce significant developments against each of these objectives over the course of the next 60 days.
In the second quarter, e-Commerce revenue for its connectivity products and services continues to grow, driven by record-levels of recurring airtime contracts and elevated hardware sales. The Company continues to see near-term growth opportunities in this business segment as it secures new connectivity contracts and expands its mission-critical communications capabilities for enterprise, government, and humanitarian organizations across Europe. Its global reach and increased focus on higher margin airtime services drove recurring airtime revenue and billable customers to record levels.
Growth in the Company's sales of OPKO Healthcare ("OPKO")-branded human health and wellness products on Alibaba Group Holding Limited's ("Alibaba") Tmall Global in China continues despite limited permissible inventory levels. The Company is working to secure increased inventory volumes to meet demand while it awaits regulatory approval to introduce OPKO pet health products as interest remains high. Approval of the sale of OPKO pet health products is still expected during the fourth quarter with sales to commence approximately 12 weeks after receipt. All current OPKO products are not subject to any tariffs as they are not produced in the United States.
As disclosed on April 11, 2025, the Company is strategically adjusting its e-Commerce development program in light of evolving trade conditions between the United States and China. While the planned launch of our Florida Sunshine-branded line of US-produced vitamins and related products in China is temporarily on hold, we are pleased to announce that Florida Sunshine products are now launching in the UK and EU. We remain committed to bringing these products to the Chinese market when conditions are more favorable.
Second Quarter 2025 Conference Call Notification
NextPlat's Interim CEO, President and CEO of Global Operations, David Phipps, and Chief Financial Officer, Cecile Munnik, will host a conference call at 8:30 a.m. Eastern today to discuss the results for the second quarter ended June 30, 2025, and recent developments.
To access the call, please use the following information:
Please call the conference telephone number 5-10 minutes prior to the start time. An operator will register your name and organization.
The conference call will be broadcast live and available for replay at and via the investor relations section of the Company's website at . A replay of the conference call will be available after 12:00 p.m. Eastern time through August 21, 2025.
The financial information included in this press release should be read in conjunction with the Company's Quarterly Report on Form 10-Q for the quarter ended June 30, 2025 filed with the Securities and Exchange Commission.
About NextPlat Corp
Nextplat is a global consumer products and services company providing healthcare and technology solutions through e-Commerce and retail channels worldwide. Through acquisitions, joint ventures and collaborations, the Company seeks to assist businesses in selling their goods online, domestically, and internationally, allowing customers and partners to optimize their e-Commerce presence and revenue. NextPlat currently operates an e-Commerce communications division offering voice, data, tracking, and IoT products and services worldwide as well as pharmacy and healthcare data management services in the United States through its subsidiary, Progressive Care.
Forward-Looking Statements
Certain statements in this release constitute forward-looking statements. These statements include the capabilities and success of the Company's business and any of its products, services or solutions. The words "believe," "forecast," "project," "intend," "expect," "plan," "should," "would," and similar expressions and all statements, which are not historical facts, are intended to identify forward-looking statements. These forward-looking statements involve and are subject to known and unknown risks, uncertainties and other factors, including the Company's ability to launch additional e-commerce capabilities for consumer and healthcare products and its ability to grow and expand as intended, any of which could cause the Company to not achieve some or all of its goals or the Company's previously reported actual results, performance (finance or operating), including those expressed or implied by such forward-looking statements. More detailed information about the Company and the risk factors that may affect the realization of forward-looking statements is set forth in the Company's filings with the Securities and Exchange Commission (the "SEC"), copies of which may be obtained from the SEC's website at . The Company assumes no, and hereby disclaims any, obligation to update the forward-looking statements contained in this press release.
Media and Investor Contact for NextPlat Corp:
Michael Glickman
MWGCO, Inc.
917-397-2272
[email protected]
SOURCE NextPlat Corp.
WANT YOUR COMPANY'S NEWS FEATURED ON PRNEWSWIRE.COM?
440k+
Newsrooms &
Influencers
9k+
Digital Media
Outlets
270k+
Journalists
Opted In
GET STARTED
Financial StatementExecutive Change
06 Aug 2025
The US Department of Health and Human Services (HHS), under the leadership of Secretary Robert F. Kennedy Jr., further distanced itself from mRNA technology on Tuesday. The health department's Biomedical Advanced Research and Development Authority (BARDA), which helps fund the development of vaccines, therapeutics and diagnostics to combat public health emergencies, is terminating about $500 million in investments across 22 mRNA vaccine programmes.Kennedy said the wind-down of agency support for the technology is because "the data show these vaccines fail to protect effectively against upper respiratory infections like COVID and flu," though he did not provide any data to support this claim. It's not the first time Kennedy has cast doubt, without scientific evidence, on the safety and efficacy of mRNA vaccines. Back in 2021, he spoke out against a proposed school vaccine mandate in Louisiana, calling the mRNA COVID-19 vaccines developed by Moderna, and partners Pfizer and BioNTech, "the deadliest vaccine ever made."In an interview with FirstWord earlier this year, virology and immunology expert Paul Offit called Kenendy a "science denialist and a virulent anti-vaccine activist" (see – ViewPoints: A vaccine expert's grim verdict on Kennedy's health agenda).After assuming leadership of HHS, Kennedy directed the Centers for Disease Control and Prevention to no longer recommend COVID-19 vaccines for pregnant women and healthy children between the ages of six months and 17 years old. This change is now being challenged in court by a coalition of healthcare organisations, including the American Academy of Pediatrics, who called it a "baseless and uninformed policy decision" that exposes "vulnerable populations to a serious illness with potentially irreversible long-term effects and, in some cases, death."The broad grant terminations come a few months after Moderna disclosed that BARDA was pulling its funding for an experimental mRNA-based bird flu jab. At the time, an HHS spokesperson claimed that mRNA technology was "under-tested," and that it was "not scientifically or ethically justifiable" to continue funding Moderna's H5N1 mRNA vaccine.After that announcement, Pfizer CEO Albert Bourla came to the defense of Moderna, calling the HHS's claims "completely inaccurate."Moderna is also developing an mRNA based flu vaccine, which it plans to incorporate with its next-generation COVID-19 jab for a single combination shot to protect against both viral illnesses. While the biotech has shared compelling data on both components, this latest move from Kennedy casts doubt on the combination vaccine's regulatory chances.Pharma caught in the falloutThe mRNA vaccine grant terminations will also affect Emory University and Tiba Biotech, who have had their contracts cancelled, and BARDA's mRNA-related work with Luminary Labs, ModeX and CSL Seqirus will be narrowed.Proposals or pre-award solicitations from Pfizer, Sanofi, CSL Seqirus and Gritstone were also rejected, and a nucleic acid-based vaccine project with AstraZeneca will be restructured. For future contracts, HHS will prioritise whole-virus vaccine technologies and other, unspecified "novel platforms."
VaccinemRNAExecutive Change
05 Aug 2025
Department of Health and Human Services Secretary Robert F. Kennedy Jr. has targeted vaccines extensively since taking office as HHS secretary.\n The Department of Health and Human Services (HHS) has announced a plan to end mRNA vaccine work funded by the Biomedical Advanced Research and Development Authority (BARDA), a major escalation of Secretary Robert F. Kennedy Jr.’s campaign against both vaccines and mRNA in general.While no timeline for the wind-down was given, the decision affects 22 projects valued at around $500 million collectively, according to an Aug. 5 release. Additionally, no new mRNA projects will be started.“We reviewed the science, listened to the experts and acted,” RFK Jr. said in the release. “BARDA is terminating 22 mRNA vaccine development investments because the data show these vaccines fail to protect effectively against upper respiratory infections like COVID and flu. We’re shifting that funding toward safer, broader vaccine platforms that remain effective even as viruses mutate.”The cancellation ends the contract for Moderna’s bird flu vaccine candidate, which the company already announced was being cut by the HHS in May. The Big Biotech isn\'t aware of any new contract cancellations with BARDA and doesn\'t have any active collaborations with the unit, a Moderna spokesperson confirmed with Fierce.The move also terminates contracts to Emory University and Tiba Biotech and reduces mRNA work from contracts with Luminary Labs, ModeX and Seqirus.In addition, proposals from Pfizer, Sanofi and others that involve mRNA have been rejected, and AstraZeneca’s nucleic acid vaccine collaboration with the Department of Defense will be restructured, with the HHS’ contributions immediately canceled, an HHS spokesperson told Fierce Biotech on Aug. 5. BARDA contracts that are nearing completion, like Arcturus’ for a bird flu mRNA vaccine and Amplitude’s for a trans-amplifying RNA therapeutic platform, will be allowed to finish, according to the release.The HHS has also told Global Health Investment Corporation—which manages BARDA’s investment fund, BARDA Ventures—to “cease all mRNA-based equity investments.”RFK Jr. said that instead of mRNA, the agency would invest in “better solutions,” like vaccines that use whole viruses and undisclosed novel platforms. Asked for specifics, the HHS spokesperson pointed Fierce to the May 1 announcement of Generation Gold Standard, a planned universal vaccine platform supported by the HHS and the National Institutes of Health (NIH). “The long-term effects and safety of these vaccines have not necessarily been studied,” the HHS spokesperson said. “Until we can prove that mRNA technology is safe and effective, we have to focus on other platforms.”The spokesperson cited a recent Senate report accusing the Biden administration of “efforts to downplay and delay warning the public about the risks of myocarditis associated with the mRNA COVID-19 vaccines.” Cases of myocarditis, or inflammation of the heart muscle, have been observed after mRNA COVID vaccination in a few rare cases. Catching COVID-19 increases the risk of myocarditis at a higher rate than vaccination does.The lack of long-term safety data on mRNA vaccines is because the first products to receive approval were for use against COVID-19. Multiple studies conducted so far have concluded that the vaccines are safe and effective, and the research underlying the science goes back to the discovery of mRNA in the early 1960s.“The announcement that HHS will discontinue mRNA vaccine development funding through BARDA represents a concerning policy shift that could undermine America\'s leadership in critical biotechnology innovation,\" Jeff Coller, Ph.D., Bloomberg Distinguished Professor of RNA biology and therapeutics at Johns Hopkins University, said in an Aug. 6 statement to Fierce. \"The science is clear: mRNA medicines are safe and highly effective.\"\"This decision risks ceding America\'s competitive advantage in biotechnology to other nations that are expanding their investments in mRNA research and development,\" Coller added. \"As other countries advance these proven, safe and effective therapies, American patients may increasingly depend on foreign innovation for breakthrough treatments.\"mRNA is a type of molecule that occurs in every living cell. mRNA vaccines work by instructing the body to make proteins that are similar to those found on pathogens. Once created, the immune system can respond to those proteins and learn to recognize them in the event of a future infection. The mRNA message itself is quickly degraded by the body.Moderna’s mRNA vaccine for COVID-19 was developed in partnership with the first Trump administration as part of Project Warp Speed. RFK Jr. has made a career out of anti-vaccine conspiracy and has targeted vaccines extensively since taking office as HHS secretary. In June, he removed all 17 members of the Centers for Disease Control and Prevention\'s (CDC\'s) vaccine advisory panel, replacing them with eight new members well-known for being critical of vaccines and COVID-19 measures.Those eight new CDC advisory panel members have much less experience in vaccine research than their predecessors, with four of them never publishing scientific research on vaccines during their careers, according to an analysis by Science.And, under RFK Jr., the NIH has allegedly ended dozens of grants funding research on vaccine hesitancy. NIH officials also told scientists to remove references to mRNA vaccines from grant applications in March, according to a report from KFF Health News.Editor\'s note: This story was updated at 10 a.m. ET on Aug. 6 with a statement from Jeff Coller, Ph.D.
VaccinemRNAExecutive Change
100 Deals associated with OPKO Health, Inc.
Login to view more data
100 Translational Medicine associated with OPKO Health, Inc.
Login to view more data
Corporation Tree
Boost your research with our corporation tree data.
login
or
Pipeline
Pipeline Snapshot as of 04 Dec 2025
The statistics for drugs in the Pipeline is the current organization and its subsidiaries are counted as organizations,Early Phase 1 is incorporated into Phase 1, Phase 1/2 is incorporated into phase 2, and phase 2/3 is incorporated into phase 3
Discovery
7
8
Preclinical
Phase 1
2
2
Phase 2
Phase 3
3
1
Approved
Other
41
Login to view more data
Current Projects
| Drug(Targets) | Indications | Global Highest Phase |
|---|---|---|
Calcifediol ( VDR ) | Hyperparathyroidism, Secondary More | Approved |
Somatrogon(OPKO Health, Inc.) ( GHR ) | Growth hormone deficiency More | Phase 3 |
NT-002 | Hyperparathyroidism, Secondary More | Phase 3 |
MDX-2001 ( CD28 x CD3 x Trop-2 x c-Met ) | Advanced Malignant Solid Neoplasm More | Phase 1/2 |
OPK-0018 | Asthma More | Phase 1 |
Login to view more data
Deal
Boost your decision using our deal data.
login
or
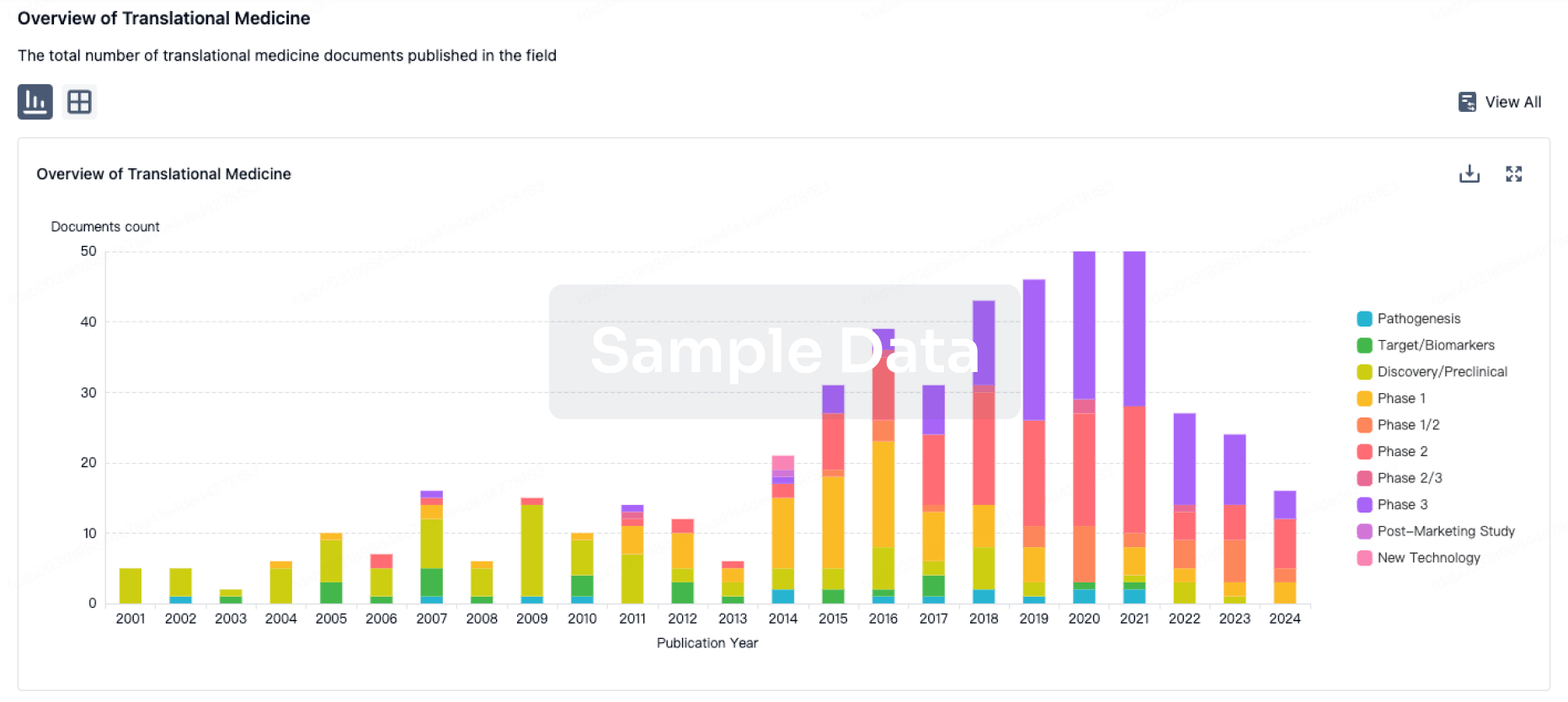
Translational Medicine
Boost your research with our translational medicine data.
login
or
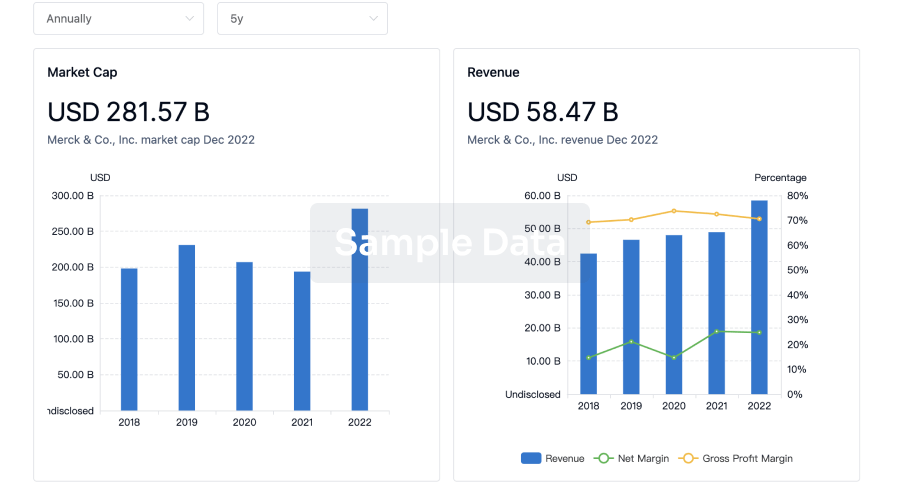
Profit
Explore the financial positions of over 360K organizations with Synapse.
login
or
Grant & Funding(NIH)
Access more than 2 million grant and funding information to elevate your research journey.
login
or
Investment
Gain insights on the latest company investments from start-ups to established corporations.
login
or
Financing
Unearth financing trends to validate and advance investment opportunities.
login
or
AI Agents Built for Biopharma Breakthroughs
Accelerate discovery. Empower decisions. Transform outcomes.
Get started for free today!
Accelerate Strategic R&D decision making with Synapse, PatSnap’s AI-powered Connected Innovation Intelligence Platform Built for Life Sciences Professionals.
Start your data trial now!
Synapse data is also accessible to external entities via APIs or data packages. Empower better decisions with the latest in pharmaceutical intelligence.
Bio
Bio Sequences Search & Analysis
Sign up for free
Chemical
Chemical Structures Search & Analysis
Sign up for free


